Wageningen University & Research | FEM-31806 | Models for Ecological Systems | FEM | PPS | WEC
Template 1 - ODE basics in R
This is a code template that you can use as backbone for an R script that implements an ODE model into code in order to solve it numerically. Additional template files that will be supplied later will expand upon this template.
The template consists of (at least) 4 steps:
- defining a function to create a named vector with initial state values
- defining a function to create a named vector with parameter values
- defining a function that returns the rates of change of the state variables with respect to time
- solving the model numerically using the
odefunction in thedeSolvepackage.
Here, we use the convention that we give a model an informative name, here simply Modelname, so that we can name the functions in steps 1-3 as listed above the names InitModelname, ParmsModelname, RatesModelname, respectively (thus, we can replace “Modelname” in the name of these 3 functions with the name of the model we are working on).
Code template
A file containing the code template as shown in the code chunk below
can be downloaded
here. Places
indicated by ... need to be filled out (other parts of the
template consist of explanation, and include placeholders for parts that
will be completed in later lectures and templates):
# Template 1 - ODE basics in R
# See: https://wec.wur.nl/mes for explanation and examples
# Load the deSolve library
library(deSolve)
# Define a function that returns a named vector with initial state values
InitModelname <- function(... = ...) {
# Gather function arguments in a named vector
y <- c(... = ...)
# Return the named vector
return(y)
}
# Define a function that returns a named vector with parameter values
ParmsModelname <- function(... = ..., # explain the parameter
... = ... # explain the parameter
) {
# Gather function arguments in a named vector
y <- c(... = ...,
... = ...)
# Return the named vector
return(y)
}
# Define a function for the rates of change of the state variables with respect to time
RatesModelname <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)),{
# Optional: forcing functions: get value of the driving variables at time t (if any)
# see template 2
# Optional: auxiliary equations
# Rate equations
...
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(...)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
# see template 4
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- NULL
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(c(RATES, # the rates of change of the state variables (same order as 'y'!)
MB), # the rates of change of the mass balance terms (or NULL)
AUX) # optional additional output per time step
return(outList)
}
)
}
# Numerically solve the model using the ode() function
states <- ode(y = InitModelname(...),
times = seq(from = ..., to = ..., by = ...),
func = RatesModelname,
parms = ParmsModelname(...),
method = ...)
# Plot
plot(states)An example with explanation
To show how the template can be used to implement an ODE model in R, let’s model a population with logistic growth:
where is the intrinsic population growth rate and is the carrying capacity, with:
Let’s assume that the initial size of the population (thus at time ) is .
Initial state variables
Ideally, we define a function, let’s call it
InitLogistic, that returns a named vector with
initial state values:
InitLogistic <- function(N = 5) {
# Gather function arguments in a named vector
y <- c(N = N)
# Return the named vector
return(y)
}After defining the function, we can retrieve the initial state values (at their defaults), simply using:
InitLogistic()## N
## 5But, we can also retrieve the initial states while changing values compared to their defaults, e.g.:
InitLogistic(N = 6)## N
## 6The here-described way of defining a named vector with initial state values via first defining a function and then calling the function may at first seem like an inefficient method. However, when models become more complex (as they often are), this extra effort easily pays off during simulation and scenario testing.
Parameter values
In a similar way, we can define a function, let’s call it
ParmsLogistic, that returns a named vector with
parameter values. Parameters and their default values can be specified
in the function arguments, so that we can call the function without
having to specify their value every time, yet still have the possibility
to give them a specific value. For some parameters, it may be sufficient
to specify them within the function body, e.g., when there are
parameters that will have a fixed value throughout all model analyses
and are thus not going to be changed.
ParmsLogistic <- function(r = 0.3, # intrinsic population growth rate
K = 100 # carrying capacity
) {
# Gather function arguments in a named vector
y <- c(r = r,
K = K)
# Return the named vector
return(y)
}Now, in a similar fashion as above, we can retrieve the parameters, at their default values, simply using:
ParmsLogistic()## r K
## 0.3 100.0Alternatively, we can retrieve the parameters at their default values while changing the value of a/some specified parameter(s) simply by passing the value to the respective function argument(s):
ParmsLogistic(r = 0.5)## r K
## 0.5 100.0Rates of change
Now we can specify a function that calculates the rates of change of
the state variables with respect to time. Let’s call the function
RatesLogistic. Check the documentation of the
ode function, ?ode(), to see the requirements
that the function RatesLogistic should adhere to. Notably,
the inputs to the function should be: t, y,
parms, in that order (although we can call them
differently)! Also, pay attention to the way the function should return
its output! As the help-file of the ode function states,
the data returned should be a list, where the first element is
a vector with the rates of change (in the same order as
y!), and all other elements are (optional) extra
output, which should be named.
In the example below, we let the function return the rate of change
of (thus ) in 2 places: both in the first
element of the returned list (thus used by the ode function
for the numerical integration), but also in the optional extra
list elements (which should be named and which will be included in the
output for each specific time point). By doing so, we not only
numerically integrate the dynamics of population , but also keep track of the rate of change
of at the specified time points.
RatesLogistic <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)),{
# Optional: forcing functions: get value of the driving variables at time t (if any)
# see template 2
# Optional: auxiliary equations
# Rate equations
dNdt <- r*N*(1-N/K)
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(dNdt = dNdt)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
# see template 4
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
# here we pass the rate of change of N with respect to time back as auxiliary variable
AUX <- RATES
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(c(RATES, # the rates of change of the state variables
MB), # the rates of change of the mass balance terms if MB is not NULL
AUX) # optional additional output per time step
return(outList)
}
)
}Solving the model
After installing and loading the deSolve, we can solve
the model numerically using the ode() function. We can
check the use of this function using ?ode(), where we see
that there are (at least) 4 function arguments that need our input:
y: the initial state values, a vector. If it has named elements, these names will be used to label the output matrix;times: time sequence for which output is wanted; the first value oftimesis the initial time;func: a function that computes the values of the derivatives at time t,parms: parameters of the model passed tofunc.
Input for the function argument y can be generated using
the above-defined function InitLogistic. Likewise, input
for the function argument parms can be generated using the
above-defined function ParmsLogistic. Input for the
function argument times still needs to be defined, here
let’s solve the model for times from 0 (the initial time) to 50 with
increments of 1. For this we can simply use the short-hand code
0:50, but also the more versatile code
seq(from=0, to=50, by=1) (more versatile because we can
easily choose another time-step!). Input for the function argument
func is the function RatesLogistic as defined
above. Note that we pass the entire function RatesLogistic
to this argument, and do not evaluate the function, thus we do not pass
RatesLogistic() as input to the func argument,
but RatesLogistic! Below, we also pass a value to the
method argument to specify that we want to use an adaptive
stepsize Runge-Kutta integration method: “ode45”).
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 50, by = 1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "ode45")Exploring the solved model
The object states now contains the solved model: it is a
matrix of class deSolve (see the
Value slot in the ?ode() function
documentation):
class(states)## [1] "deSolve" "matrix"To explore the object states, we can thus look at the
top (header; using the head function) and bottom (tail;
using the tail function) rows of the matrix, and use the
defaults plotting method (using the function plot) to plot
the solved model:
head(states)## time N dNdt
## [1,] 0 5.000000 1.425000
## [2,] 1 6.633260 1.857977
## [3,] 2 8.750883 2.395531
## [4,] 3 11.461553 3.044364
## [5,] 4 14.875003 3.798704
## [6,] 5 19.085896 4.632955tail(states)## time N dNdt
## [46,] 45 99.99740 0.0007814116
## [47,] 46 99.99807 0.0005788920
## [48,] 47 99.99857 0.0004288582
## [49,] 48 99.99894 0.0003177084
## [50,] 49 99.99922 0.0002353655
## [51,] 50 99.99942 0.0001743638plot(states)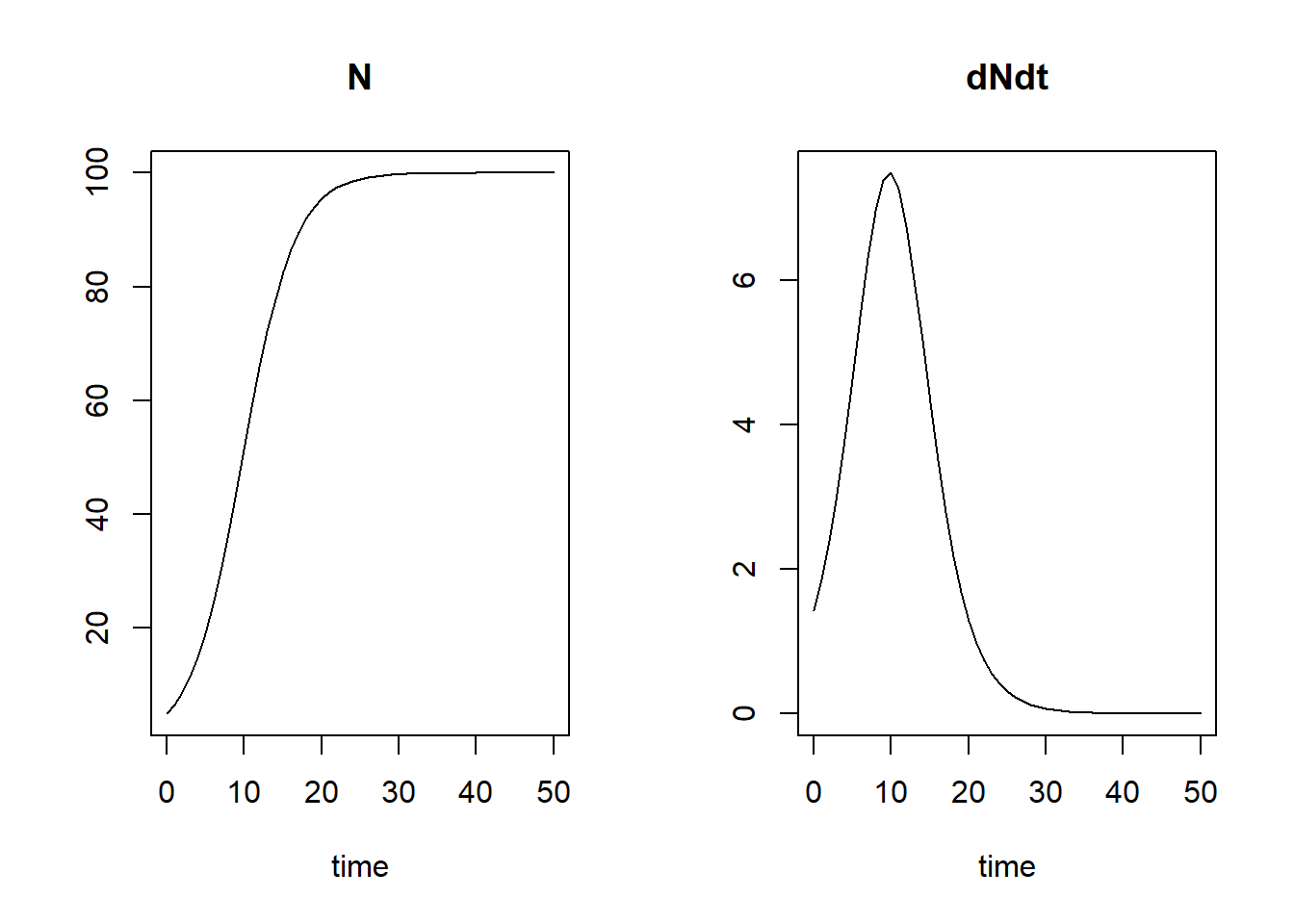
The first column of states is called “time”, with the
values corresponding to what we supplied to the function argument
times when calling the ode function. The
second column is called “N”: this is because the
InitLogistic function that we used to pass the initial
state value to the argument y returns a vector with the
name “N”, so this is the population size. The third and last column is
called “dNdt”: this is the optional auxiliary data that we
chose to also return. The default plotting method will plot how the
state variables, as well as the returned auxiliary variables, evolve
over time.
Note that at time 0, (our set
initial value!) and . But note that at time 1,
the value of is not
equal to ! Thus, the
numerical integration performed by the ode function used
integration timesteps smaller than 1 time unit, while recording the
output at the times specified in the times argument (we
used here the adaptive stepsize Runge-Kutta integration method:
“ode45”). Thus, when using a variable timestep integration methods,
input to the times argument effectively specifies the
maximum timestep that will be used in the integration. You can
check for yourself whether the value of at time 1 will be when using Euler integration
(use method="euler").
Now, suppose that we want to know the value of at time 50, we have a few options to
retrieve this. First, we can simply read out the value from the
corresponding row, e.g. using the tail function as used
above (here with the extra function argument n=1 to
retrieve only the last record):
tail(states, n = 1)## time N dNdt
## [51,] 50 99.99942 0.0001743638Alternatively, we can use the subset function:
subset(states, time == 50)## N dNdt
## 9.999942e+01 1.743638e-04Or, we could use matrix indexing using the square bracket notation
[,] to retrieve either the 51th row explicitly (i.e. the
row corresponding to time 50), or the last row of the matrix (using the
function nrow):
states[51,]## time N dNdt
## 5.000000e+01 9.999942e+01 1.743638e-04states[nrow(states),]## time N dNdt
## 5.000000e+01 9.999942e+01 1.743638e-04What we see here, also from the graph above, is that the population has almost reached a value of 100 at time 50, i.e. it is very close to its carrying capacity !
Because we returned the rate of change also as an auxiliary variable, we can now plot how varies with :
plot(states[,"N"], states[,"dNdt"], type = "l", xlab = "N", ylab = "dN/dt")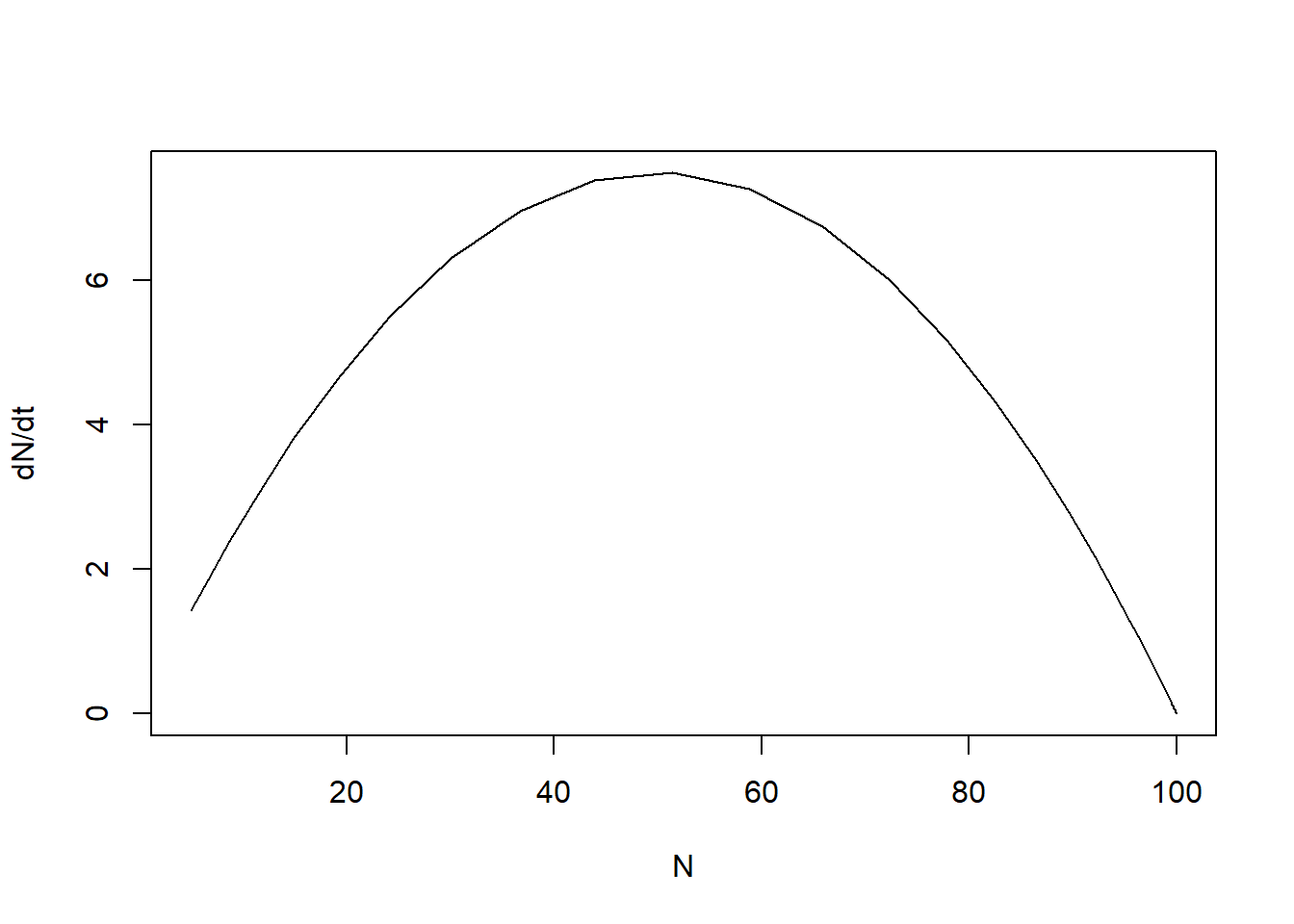
Downloadable R script
An R script with the code chunks of the above example can be downloaded here.