Wageningen University & Research | FEM-31806 | Models for Ecological Systems | FEM | PPS | WEC
Chapter 3 - Programming in R
Here, you will practice some more R skills that will be useful during the coming weeks when we define and analyse models in R:
- plotting
for-loops- writing your own functions
- using the
withfunction - working with
liststo store data
Exercise 3.1
One of the greatest strengths of R, relative to other programming
languages, is the ease with which we can create publication-quality
graphics. Here, we will try some very basic plotting in R, and do not
cover the more advanced ways of plotting using e.g. the
lattice, ggplot2 and ggvis
libraries.
iris dataset, using the
head function head(iris)## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 setosa
## 2 4.9 3.0 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5.0 3.6 1.4 0.2 setosa
## 6 5.4 3.9 1.7 0.4 setosaplot function applied to a data.frame will give
you a plot where all columns in the data.frame are plotted against each
other (i.e., pair plots). Give it a try. plot(iris)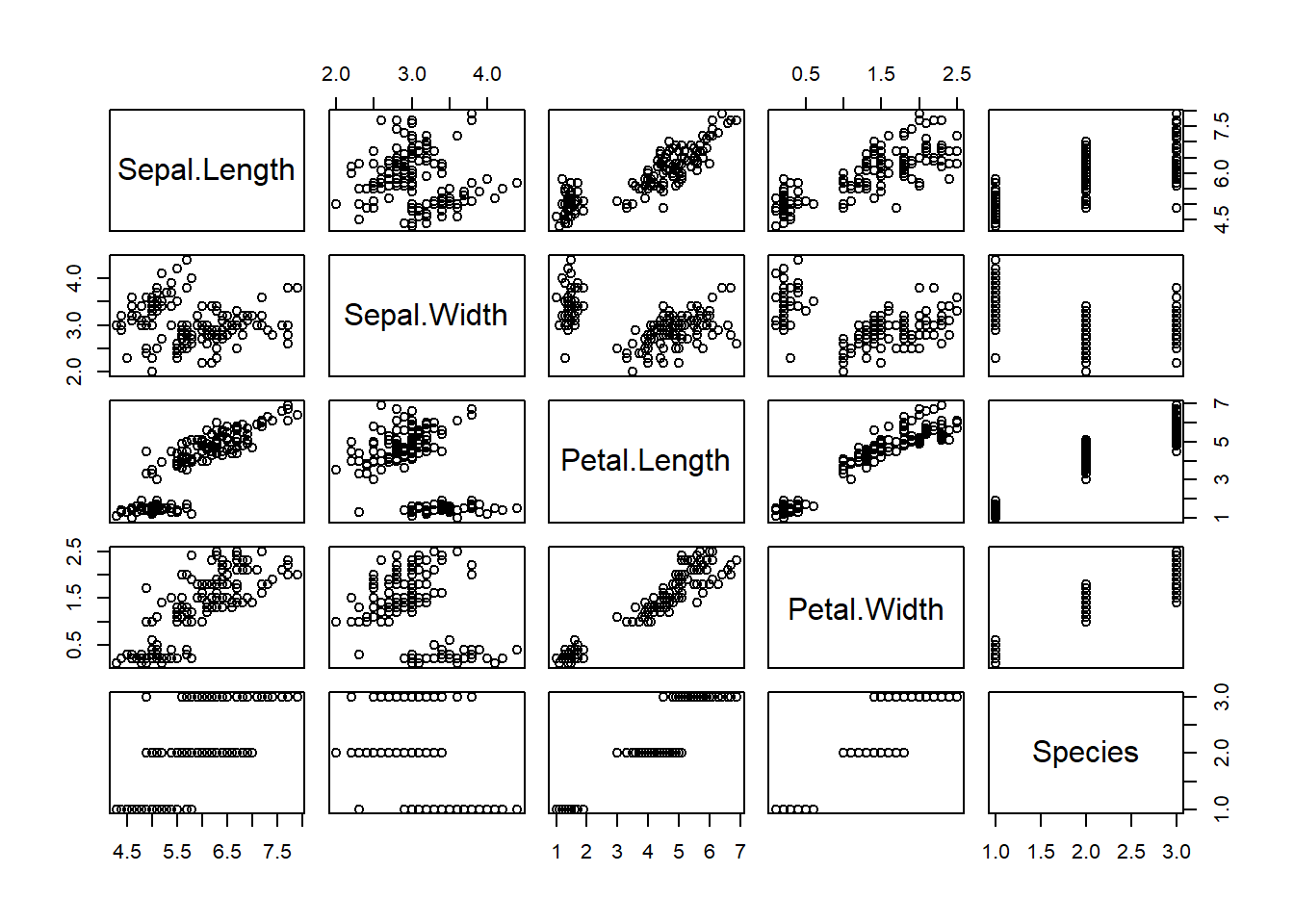
$-sign to retrieve entire
columns from a data.frame. Plot 2 specific columns against each other
the $-sign notation: plot column Sepal.Length
on the x-axis and column Sepal.Width on the y-axis.
plot(iris$Sepal.Length, iris$Sepal.Width)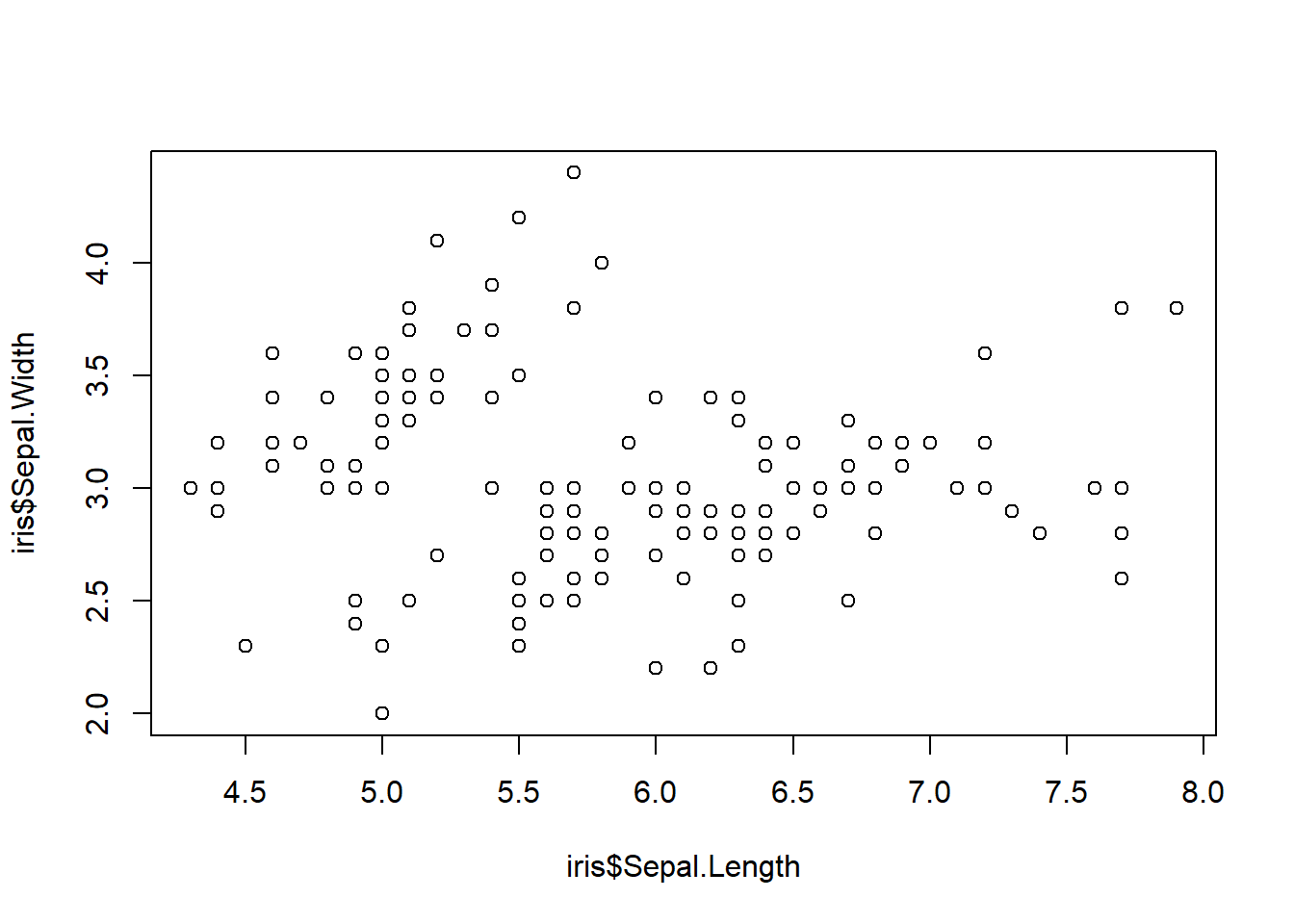
[] notation to
index specific columns from the data.frame (or
matrix). Remember that a data.frame is a 2-dimensional data
structure, and indexing the column should be done in the second
element of the index: e.g. iris[,4] returns the
4th column in the iris dataset. Create the same
plot with column Sepal.Length on the x-axis and column
Sepal.Width on the y-axis using [] notation
and the column position numbers. plot(iris[,1], iris[,2])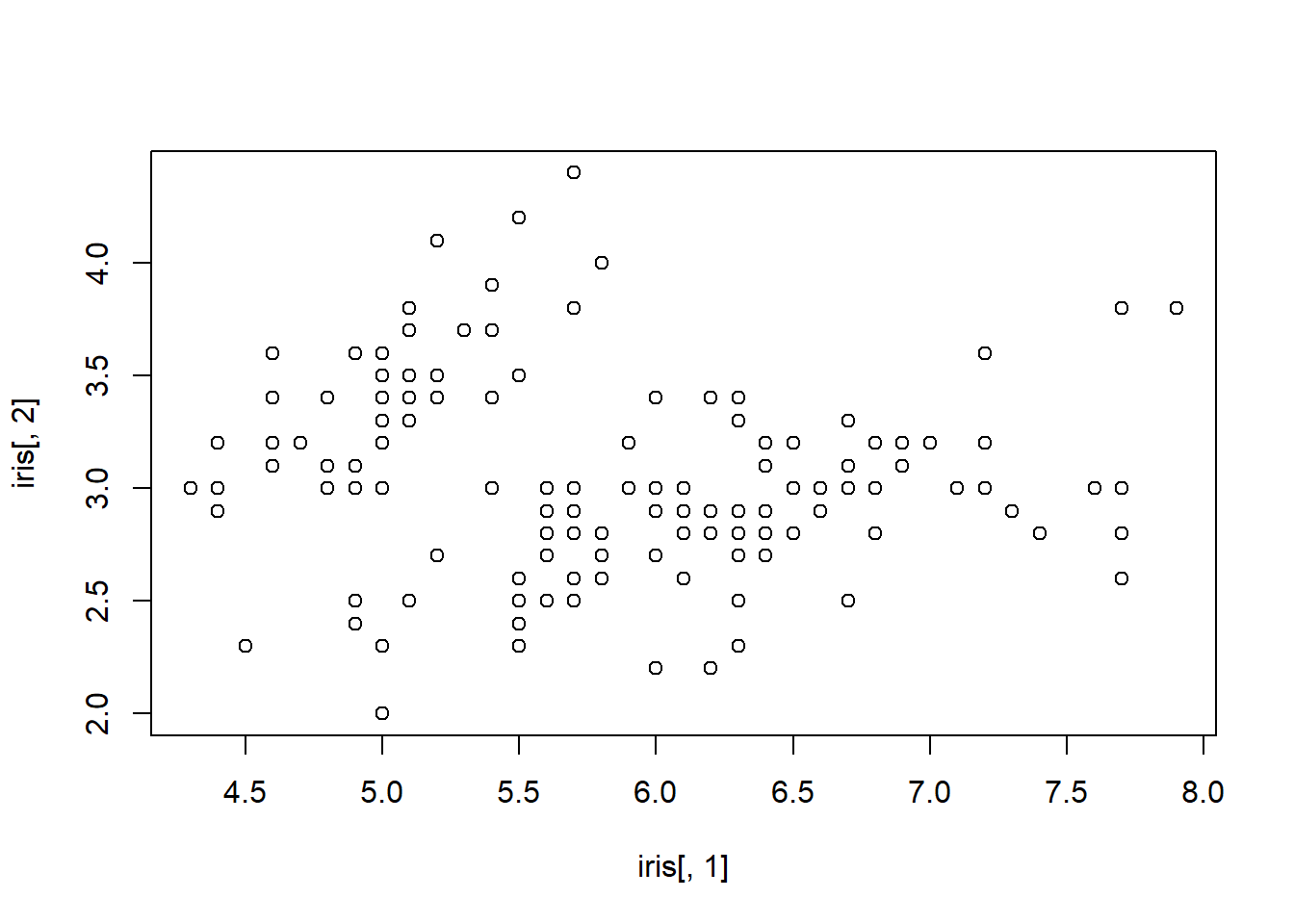
plot(iris[,"Sepal.Length"], iris[,"Sepal.Width"])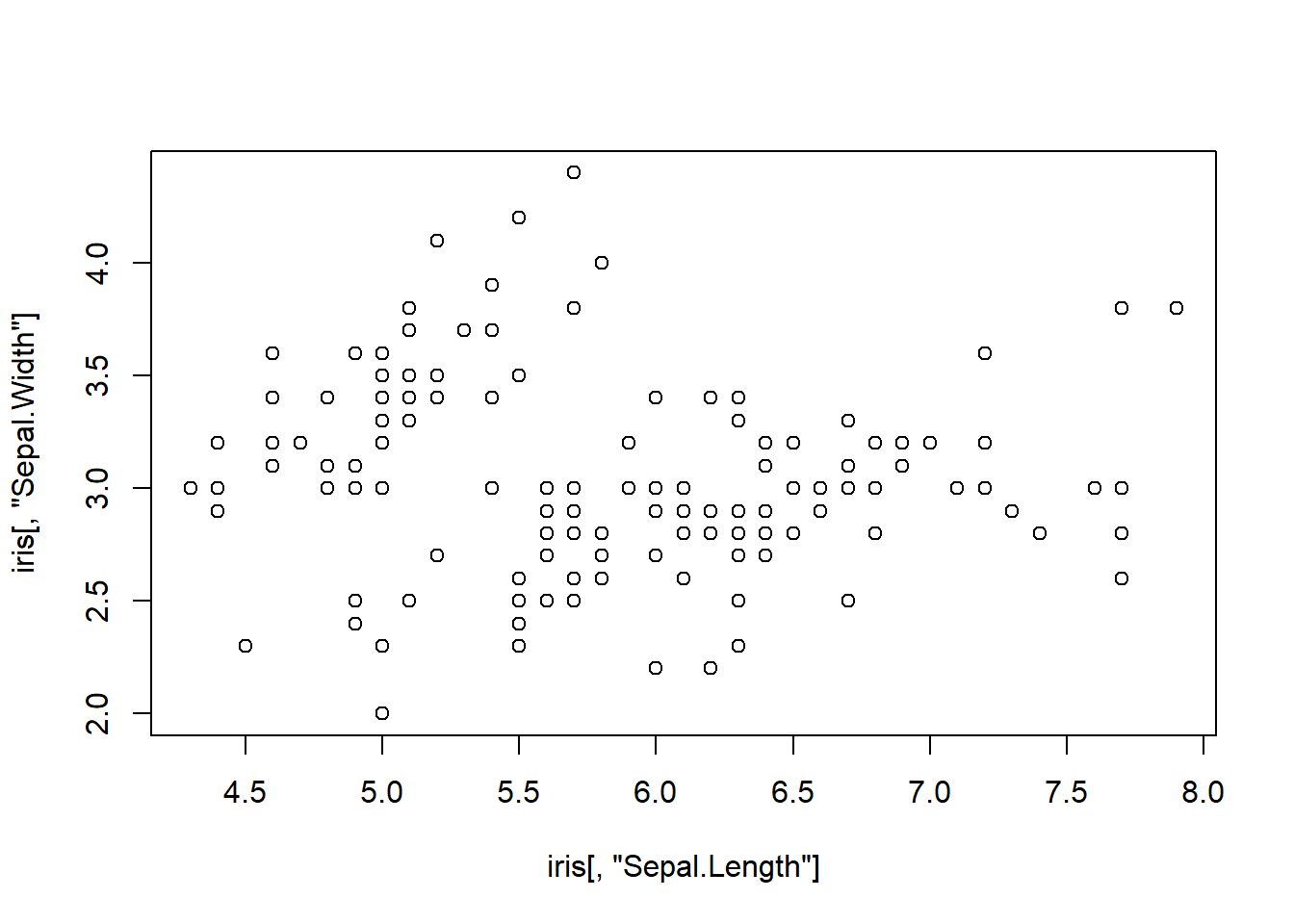
To specify the axis labels in a plot() function call,
you can use the arguments xlab and ylab for
x-axis and y-axis labels, respectively. To specify point type and
colour, use function arguements pch and col,
respectively.
plot(iris$Sepal.Length, iris$Sepal.Width,
xlab = "length", # x axis label
ylab = "width", # y axis label
pch = 16, # point type
col = "green") # point colour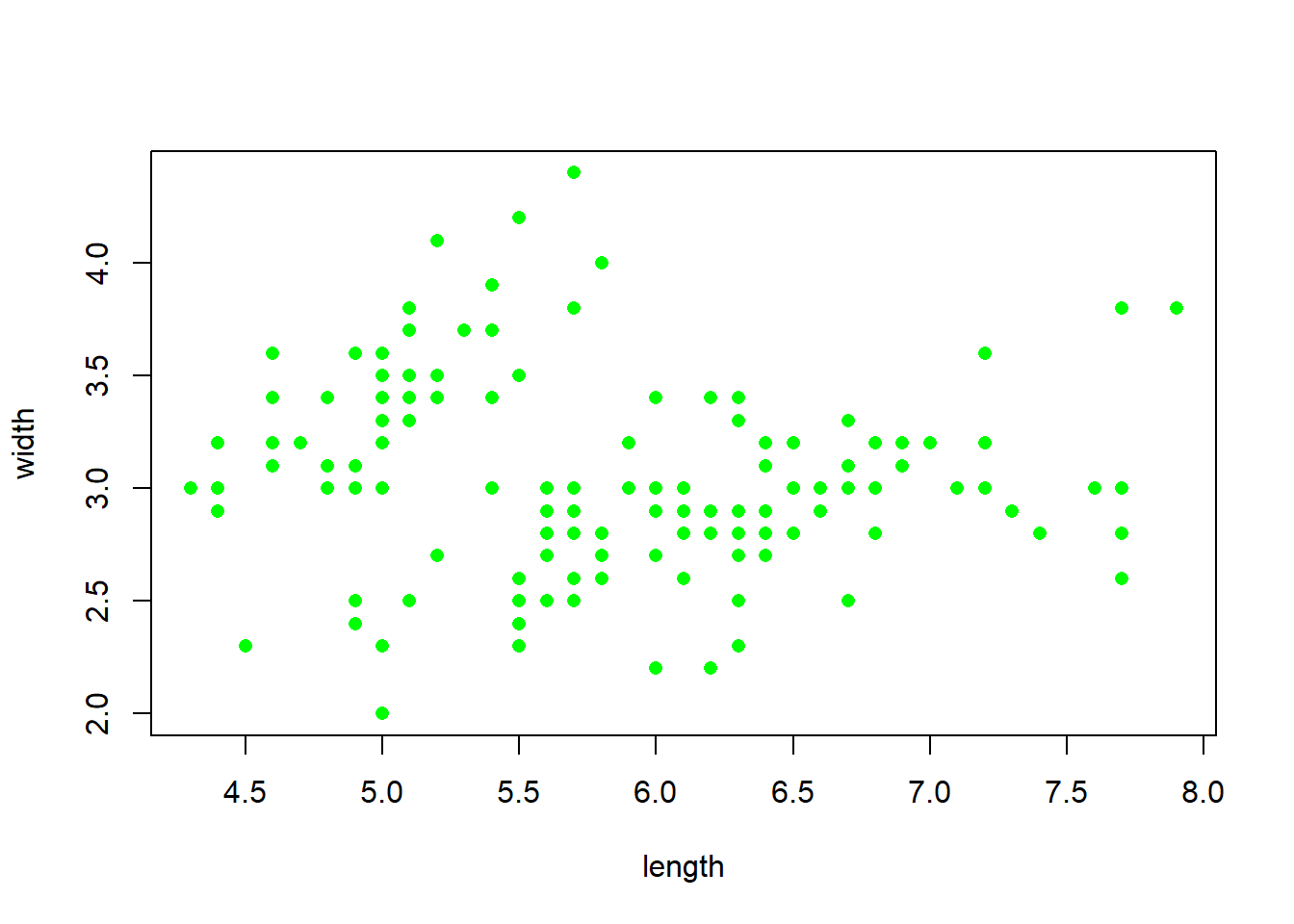
The plot function creates by default a new plot, and
using the type argument you can choose the type of plot
that you want to plot (default option is “p” which plots points, option
“l” plots a line, option “b” plots points connected by a line, and
option “n” plots only the plotting window, but no lines or points). On
top of an existing plot you can add points or lines via the functions
points and lines respectively.
Run the following code to create a data.frame with measurements of grass biomass over time:
grassBiomass <- data.frame(time = 0:50,
grass = c(8.4,8.2,5.3,6.7,7.4,12.3,15.0,13.6,9.2,10.5,12.0,
13.6,14.4,11.7,12.0,13.6,12.5,9.3,10.8,12.3,11.3,
14.9,11.2,11.2,5.1,6.4,10.9,11.2,9.8,11.5,9.0,
10.2,6.5,5.0,6.3,6.6,12.5,15.0,12.1,12.0,12.5,
12.0,12.1,13.9,11.3,9.6,11.5,8.2,9.7,10.1,8.5))time on the x-axis
and grass on the y-axis (thus use type="n"),
and set “Time” and “Grass biomass” to be the axis labels. Then, add a
black line to the plot, and after that add solid red points to the plot
(both with time on the x-axis and grass on the
y-axis). plot(grassBiomass$time, grassBiomass$grass, type = "n", xlab = "Time", ylab = "Grass biomass")
lines(grassBiomass$time, grassBiomass$grass, col = "black")
points(grassBiomass$time, grassBiomass$grass, col = "red", pch = 16)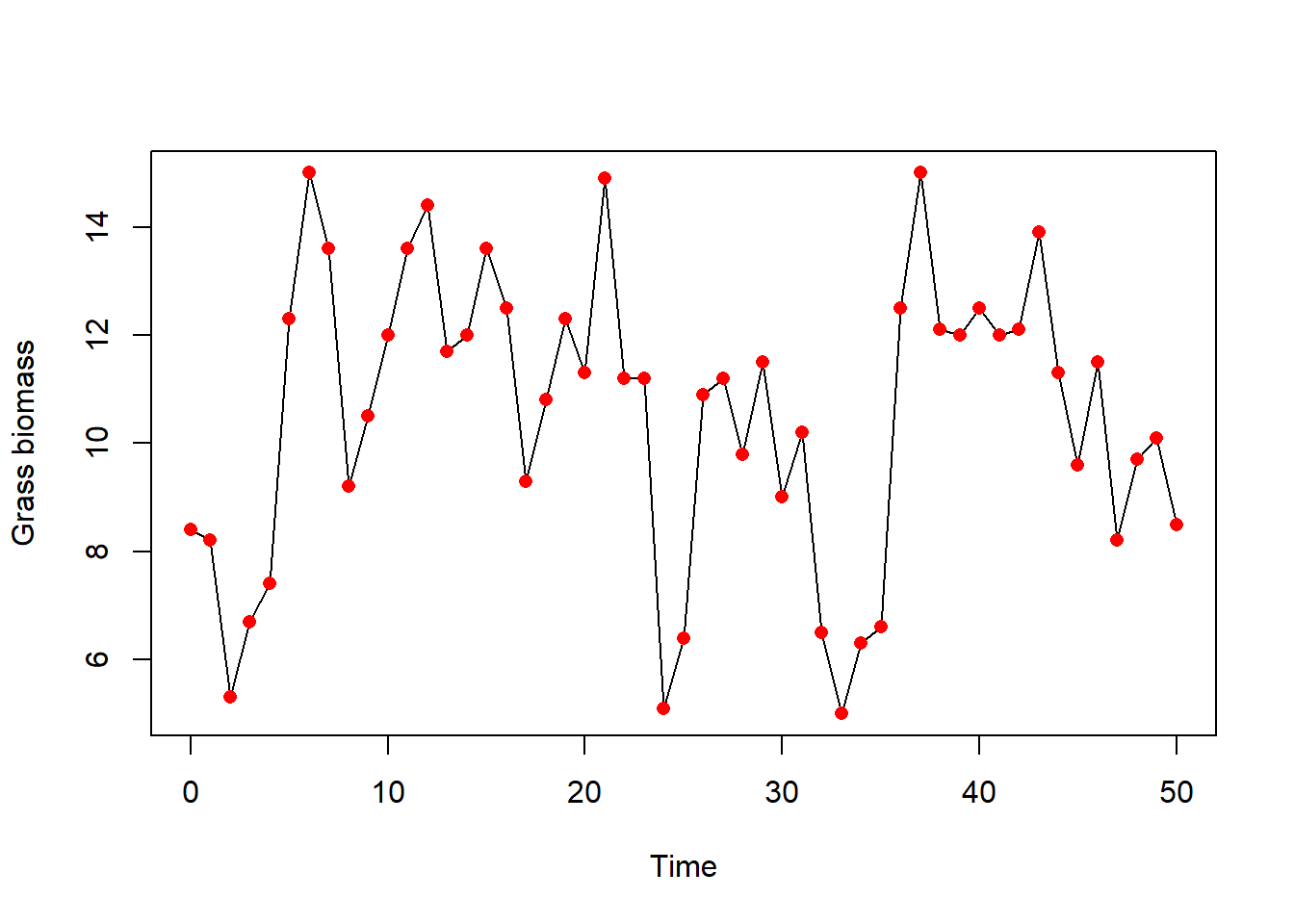
Exercise 3.2
for-loop: use the iris dataset
and print (to the console, using the print function) the
value of element on the first row of the first column, the
2nd row of the 2nd column, … etc …, up to the
5th row in the 5th column. for(i in 1:5) {
print(iris[i,i])
}## [1] 5.1
## [1] 3
## [1] 1.3
## [1] 0.2
## [1] setosa
## Levels: setosa versicolor virginica1:5 as done above, you can use
the c() function to create a sequence of values over which
the iterator i should iterate. for(i in c(1,51,101)) {
print(iris[i,])
}## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 setosa
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 51 7 3.2 4.7 1.4 versicolor
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 101 6.3 3.3 6 2.5 virginicaExercise 3.3
myFun that takes 2
function argument as input, call them x and y,
and returns an object called z, which is the product of
x and the square root of y (using function
sqrt). myFun = function(x, y) {
z <- x * sqrt(y)
return(z)
}x) and 100 (for
y): is the output of the function indeed 50? myFun(x=5, y=100)## [1] 50Exercise 3.4
Sometimes, e.g., the coming weeks when analysing models using R, you
will not supply a single value as input to a function, but rather a
named vector or list with many parameter values and/or state variables.
The with function is in such a case a very helpful way to
retrieve the elements of a named object in a concise and efficient
way.
with function: what
type of object can be passed on to the function as its first argument?
?with
# the first argument (data) may be an environment, a list, a data frame, or an integervector with element names a and
b, with values 5 and 3, respectively, and assign it to an
object called parms. parms <- c(a=5, b=3)a and b stored
in the object parms, using square bracket index notation.
parms[["a"]] + parms[["b"]]## [1] 8However, especially with objects containing many different elements,
coding in this way can become very tedious. Using the with
function, we can access the elements a and b
directly using their name (without having to index the elements using
parms[["name""]]). The with function can be
used with the following syntax:
with(data, expr)Here, data contains the elements, and expr
contains the code that is evaluated. Most conveniently, a
list is supplied to the data argument, and the
code in expr is bundled within curly brackets
{} so that you can even evaluate code that spans many
lines.
To convert the object parms into a list,
you can use the as.list function:
as.list(parms)For example, using the parms object created above, you
can use the with statement to print the values of elements
a and b to the console:
with(as.list(parms),{
# R statements here, possibly spanning many lines of code
# e.g. print the value of 'a' to the console
print(a)
# e.g. print the value of 'b' to the console
print(b)
})with function yourself: compute the
product of elements a and b without having to
index the elements using the square brackets. with(as.list(parms),{
a * b
})## [1] 15Exercise 3.5
for loop with the flexibility of a
list. You can create an empty list using
list(), and subsequently assign elements to it using double
square bracket notation [[]]. Creates an empty list called
myList, and then store each row of the iris
dataset as a separate element in the list using a for-loop.
Then, check the contents of the object myList by printing
the header to the console (function head). myList <- list()
for(i in 1:nrow(iris)) {
# store the i-th row of iris as the i-th element in the list
myList[[i]] <- iris[i,]
}
# check contents
head(myList)## [[1]]
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 setosa
##
## [[2]]
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 2 4.9 3 1.4 0.2 setosa
##
## [[3]]
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 3 4.7 3.2 1.3 0.2 setosa
##
## [[4]]
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 4 4.6 3.1 1.5 0.2 setosa
##
## [[5]]
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 5 5 3.6 1.4 0.2 setosa
##
## [[6]]
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 6 5.4 3.9 1.7 0.4 setosaSolutions as R script
Download here the code as shown on this page in a separate .r file.