Wageningen University & Research | FEM-31806 | Models for Ecological Systems | FEM | PPS | WEC
Appendix A - Events with root functions
In Template 3, we have demonstrated how you can specify discrete events that act on state variables at specified times. Although this is an easy way of implementing events, it is limited to events that act on a single state at a time, and these times need to be known at the start of a simulation!
The deSolve package allows for a more versatile specification of discrete events, where 2 different types of events are specified:
- Events that occur at known times:
- Simple changes can be specified in a data.frame (see Template 3);
- More complex events can be specified in an event function
that returns the changed values of the state variables. This function
should be defined as
function(t, y, parms, ...);
- Events occur when certain conditions are met:
- These events are triggered by a root function;
- The events themselves are specified in an event function.
Roots occur when a root function becomes zero.
By default, when a root is found, the simulation either stops (no event), or triggers an event. Thus, when a root function is defined, but not an event function, then the solver will stop the simulation at a root.
Not all solver methods (e.g. “ode45”) can handle root functions! When using root functions, use one of the solvers that can handle them, e.g. “lsoda”.
The details are explained in the ?events help page, but
let us now show a small example.
Logistic growth
Let’s use the example of logistic growth as defined in Template 1:
# Define a function that returns a named vector with initial state values
InitLogistic <- function(N = 5) {
# Gather function arguments in a named vector
y <- c(N = N)
# Return the named vector
return(y)
}
# Define a function that returns a named vector with parameter values
ParmsLogistic <- function(r = 0.3, # intrinsic population growth rate
K = 100 # carrying capacity
){
# Gather function arguments in a named vector
y <- c(r = r,
K = K)
# Return the named vector
return(y)
}
# Define a function for the rates of change of the state variables with respect to time
RatesLogistic <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)),{
# Optional: forcing functions: get value of the driving variables at time t (if any)
# see template 2
# Optional: auxiliary equations
# Rate equations
dNdt <- r*N*(1-N/K)
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(dNdt = dNdt)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
# see template 4
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
# here we pass the rate of change of N with respect to time back as auxiliary variable
AUX <- RATES
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(c(RATES, # the rates of change of the state variables
MB), # the rates of change of the mass balance terms if MB is not NULL
AUX) # optional additional output per time step
return(outList)
}
)
}After having defined these functions, we can solve the model using
the ode function and plot the results:
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 100, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "ode45")
plot(states)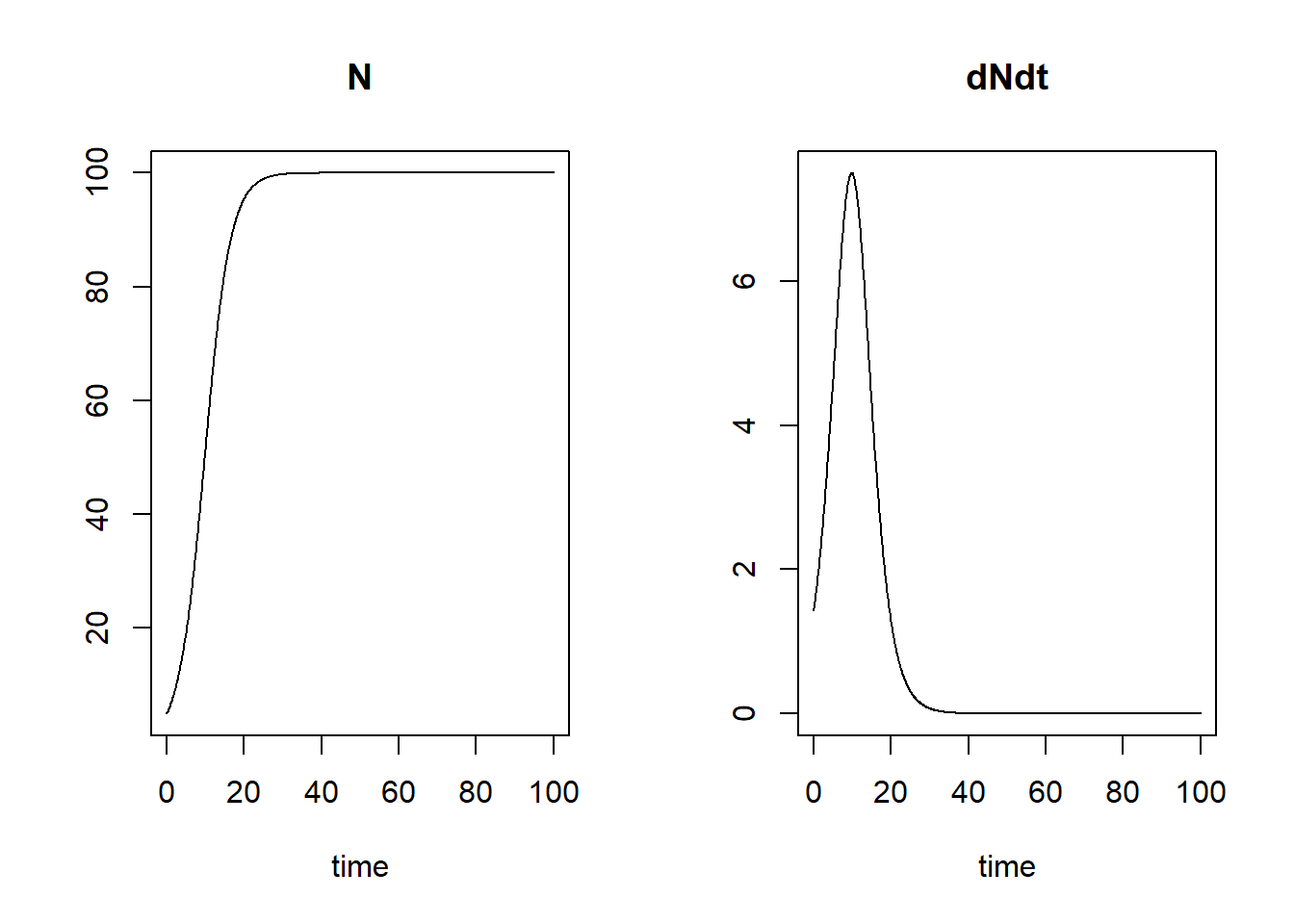
Triggering early termination
If we provide a root function but not an event
function, than the simulation will stop at a root. Therefore, let’s
create a root function, called root, that triggers
a root when the population size
reaches 99% of the carrying capacity
(i.e. a root is triggered when this function returns the value 0):
# Root function when N == 0.99*K
root <- function(t, y, parms) {
# we use the with() function in a similar way to its use in the RatesLogistic function
# so that we can easily refer to parameters and state variables by their name(s)
with(
as.list(c(t=t, y, parms)),{
r <- N - (0.99*K) # difference between N and 99% of K: thus: value 0 when N equals 99% of K
return(r) # then r == 0 this triggers a root
})
}After defining this root function, we can simulate the population and
stop the simulation when we supply the root function but do not supply
an event function. Thus, we pass object root as input to
argument rootfunc, yet the events argument
still needs input: as earlier this needs to be a list, but
now with values of elements “func” and “root”:
- in element “func” we need to specify the event function;
here we do not supply the event function because our intent is early
termination of the simulation, therefore we assign it the value
NULL; - in element “root” we need to indicate (with a logical,
value, thus either
TRUEorFALSE) whether we supply a root function to input argumentrootfunc. Here we set it toTRUEsince be are supplying a root function. However, we may not always need to supply a root function, as we can also supply an event function along with specified “time” values during which the event function will be called.
Apart from supplying the events list and root function, we set the integration method to “lsoda”, as the “ode45” method used in the templates can not handle root functions:
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 100, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "lsoda", # "ode45" does not handle root functions, but "lsoda" does
events = list(func = NULL, # NULL: therefore the simulation is stopped at a root
root = TRUE), # specifying that there is a root function
rootfunc = root) # the defined root function
plot(states)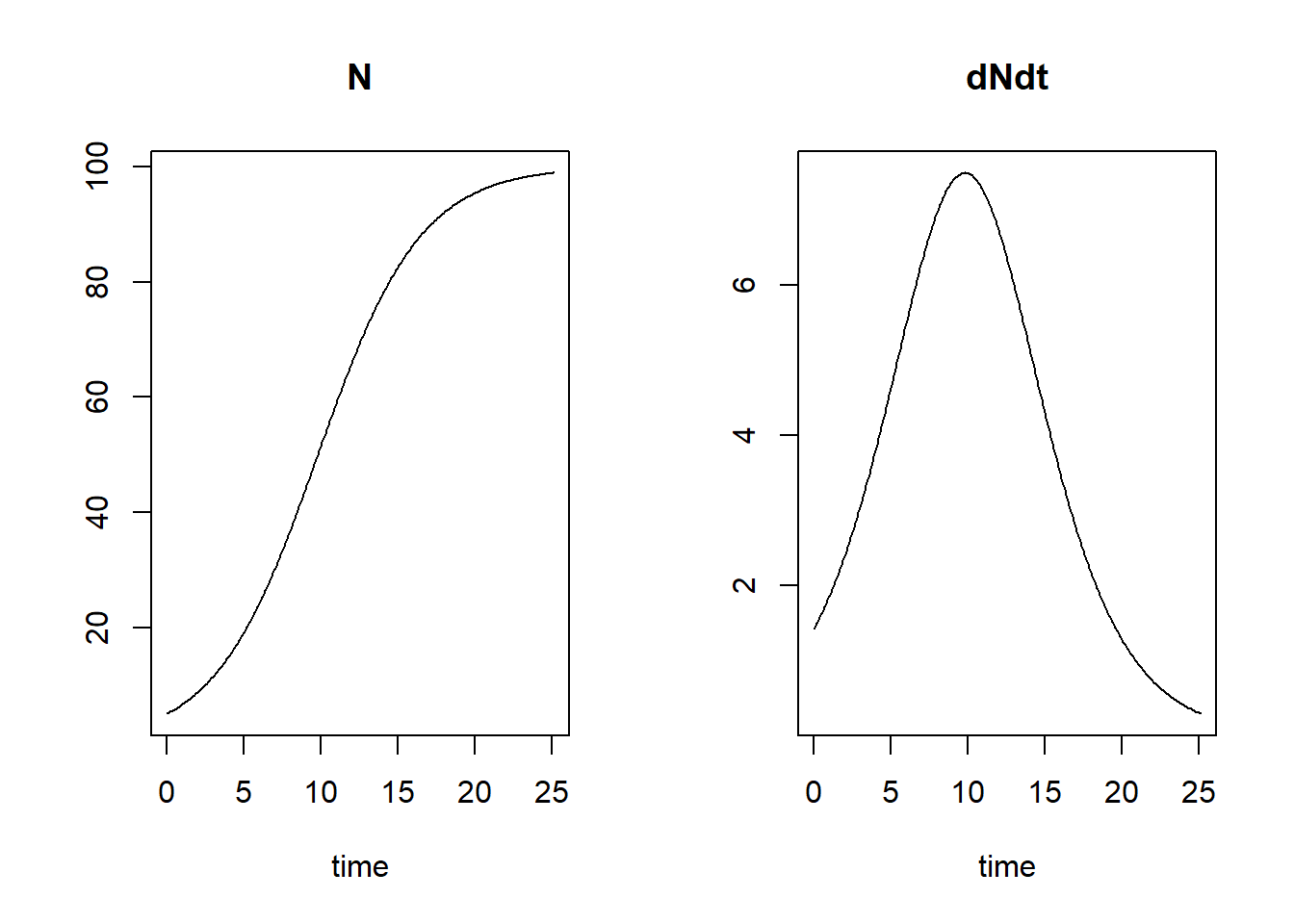
We see that the population reaches 99% of its carrying capacity around time 25, therefore the simulation is stopped and not continued till time 100.
Events at specified times
So demonstrate how we can use an event function that acts on state variables at pre-defined times, let’s simulate the population for 50 time units, and remove 25% of the population at times 10, 20, 30 and 40. For this, we need to specify an event function that removes 25% of the state variable at the specified times (thus, we replace by ), and we need to specify the times at which this event function is applied:
# specify event function
eventfun <- function(t, y, parms) {
y["N"] <- 0.75 * y["N"]
return(y)
}
# solve the model
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "ode45",
events = list(func = eventfun, # the event function, called at times in "time"
time = c(10,20,30,40), # the time at which the event function is applied
root = FALSE)) # this specifies that we do not supply a root function
# thus: the event function is called at the specified times
# plot the simulation
plot(states)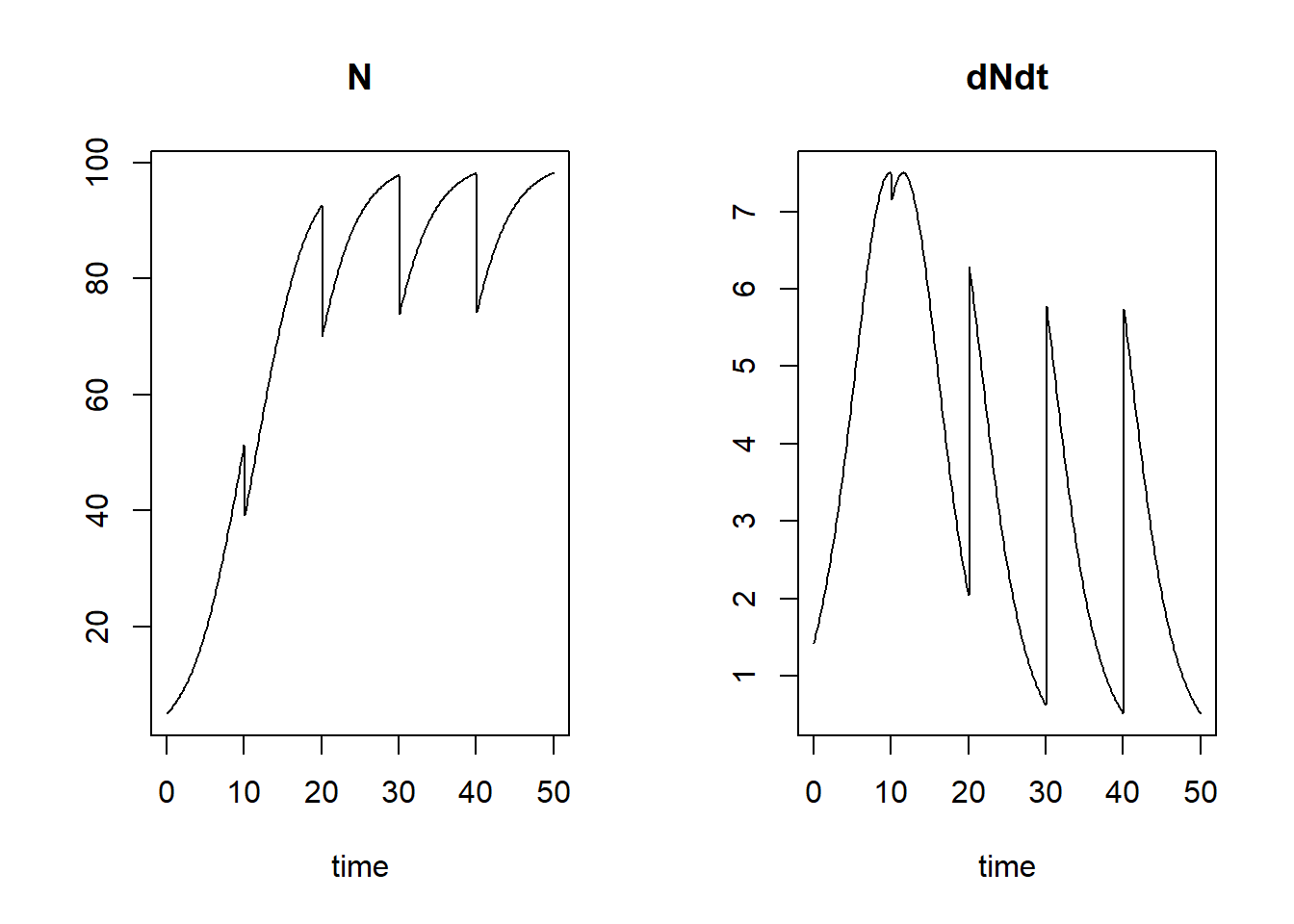
Events triggered by a root
By supplying both an event function as well as a root function, we can apply events to a state variables when some condition is met. For example, we can reduce a population by 25% when the population is large enough, e.g. when the population is at 99% of its carrying capacity :
# root function when N == 0.99*K
root <- function(t, y, parms) {
# we use the with() function in a similar way to its use in the RatesLogistic function
# so that we can easily refer to parameters and state variables by their name(s)
with(
as.list(c(t=t, y, parms)), {
r <- N - (0.99*K) # difference between N and 99% of K: thus: value 0 when N equals 99% of K
return(r) # then r == 0 this triggers a root
})
}
# specify event function
eventfun <- function(t, y, parms) {
y["N"] <- 0.75 * y["N"]
return(y)
}
# solve the model
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "lsoda",
events = list(func = eventfun, # the event function, called at times in "time"
root = TRUE), # this specifies that we do supply a root function
rootfunc = root)
# plot the simulation
plot(states)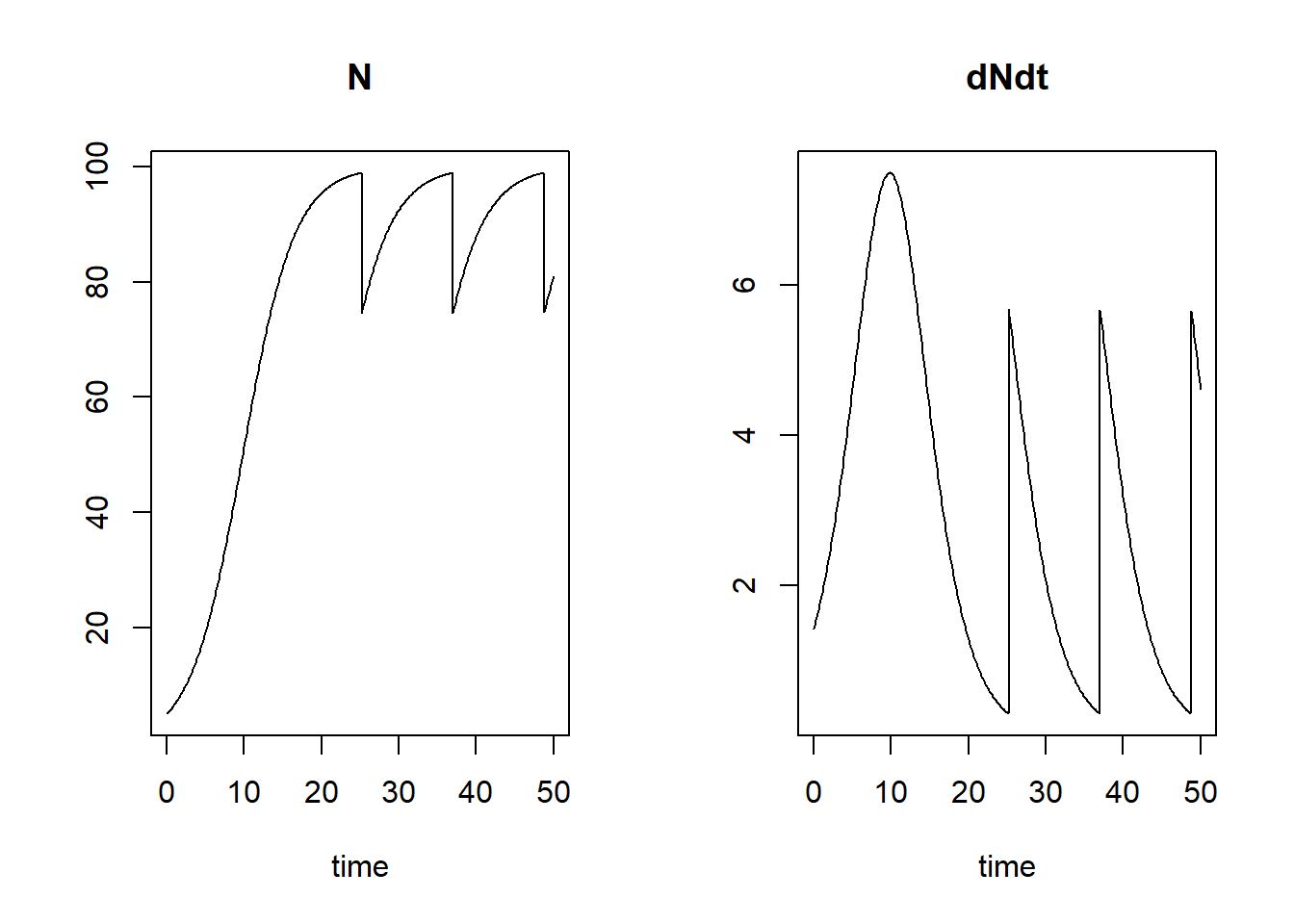
Mass balances and events
This topic is dealt with in Appendix B.
Downloadable R script
An R script with the code chunks of the above example can be downloaded here.