Wageningen University & Research | FEM-31806 | Models for Ecological Systems | FEM | PPS | WEC
Template 4 - Mass balances
This is a code template that you can use as backbone for an R script that implements a model into code in order to solve it numerically, including discontinuous events at discrete points in time.
This template only covers performing mass balance checks in the absence of discrete events. In case of performing mass balance checks in the presence of discrete events, see Appendix B.
Code template
A file containing the code template as shown in the code chunk below
can be downloaded
here. Places
indicated by ... need to be filled out (other parts of the
template consist of explanation, and include placeholders for parts that
will be completed in later lectures and templates):
# Template 4 - Mass balances
# See: https://wec.wur.nl/mes for explanation and examples
# Load the deSolve library
library(deSolve)
# Define a function that returns a named vector with initial state values
# - including the states "IN" and "OUT" to keep track of inputs and outputs
InitModelname <- function(... = ...) {
# Gather function arguments in a named vector
y <- c(... = ...,
IN = 0, # sum of inputs
OUT = 0) # sum of outputs
# Return the named vector
return(y)
}
# Define a function that returns a named vector with parameter values
# see Template 1
# Define a function for the rates of change of the state variables with respect to time
RatesModelname <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)),{
# Optional: forcing functions: get value of the driving variables at time t (if any)
# see template 2
# Optional: auxiliary equations
# Rate equations
...
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(...)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
dINdt <- ...
dOUTdt <- ...
MB <- c(dINdt = dINdt, dOUTdt = dOUTdt)
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- NULL
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(c(RATES, # the rates of change of the state variables (same order as 'y'!)
MB), # the rates of change of the mass balance terms (or NULL)
AUX) # optional additional output per time step
return(outList)
}
)
}
# We need the initial conditions later on, so store it in an object
inits <- InitModelname(...)
# Numerically solve the model using the ode() function
states <- ode(y = inits,
times = seq(from = ..., to = ..., by = ...),
func = RatesModelname,
parms = ParmsModelname(...),
method = ...)
# Plot
plot(states)
# Define a function to compute the mass balance over time
MassBalancesModelname <- function(y, ini) {
# Add "0" to the state variables' names to indicate initial condition
names(ini) = paste0(names(ini), "0")
# Using the with() function we can access elements directly by there name
# here: 'y' is a deSolve matrix
# so it is best to first convert it to a data.frame using function as.data.frame()
# and then combine it with 'ini' using the function c()
# and convert to a list using as.list()
with(as.list(c(as.data.frame(y), ini)), {
# Compute mass balance terms
BAL <- ...
# Return output BAL
return(BAL)
})
}
# Compute the mass balance and plot the result
bal <- MassBalancesModelname(y = states, ini = inits)
plot(states[,"time"], bal, type = "l", xlab = "Time", ylab = "Mass balance")An example with explanation
Let’s continue with the example of Template 1, where we modelled a population with logistic growth:
with , and . Rearrange this rate equation in order to match it with how many ecologists are using the logistic function to specify a population’s dynamics via fecundity and mortality (we now ignore that the population is subjected to emigration and immigration):
where the first term () here represents fecundity (influencing positively), whereas the second term () represents mortality (influencing negatively). Note that this is just one of the interpretations of logistic growth.
Because we now we create a new model and set of exercises, let’s call the model Logistic4, and thus names the 3 functions that we need to define: InitLogistic4, ParmsLogistic4 and RatesLogistic4. We will be defining a new function that will help us to check the mass balance of the model, so let’s call this function MassBalanceLogistic4.
Initial state variables
We first define a function that returns a named vector with
initial state values, let’s call it InitLogistic4. We can
copy InitLogistic from Template 1 - ODE basics in
R. Because it is often useful to keep track of mass balances, let’s
update the function by adding 2 state variables, “IN” and “OUT”, both
with the initial value 0, so that we can keep track of the inputs and
outputs in the system. Because at the start of a simulation both should
have a value 0, we do not need to include “IN” and “OUT” in the function
arguments, but rather can define them (with value 0) hard-coded in the
function body!
InitLogistic4 <- function(N = 5) {
# Gather function arguments in a named vector
y <- c(N = N, # population size
IN = 0, # sum of inputs
OUT = 0) # sum of outputs
# Return the named vector
return(y)
}To check, let’s run:
InitLogistic4()## N IN OUT
## 5 0 0Note that we now have specified that there are 3 state variables (called “N”, “IN” and “OUT”), the function for the rates of change of the state variables (that we will specify below) should return the rate of change of all 3 specified states!
Parameter values
Because the model parameters have not changed compared to the
function ParmsLogistic in Template 1, let’s copy that
function and call it ParmsLogistic4.
ParmsLogistic4 <- function(r = 0.3, # intrinsic population growth rate
K = 100 # carrying capacity
) {
# Gather function arguments in a named vector
y <- c(r = r,
K = K)
# Return the named vector
return(y)
}Rates of change
We can adapt the function RatesLogistic given in
Template 1 to add some lines of code to keep track of the in and out
flow so that we can compute mass balances, let’s call the new function
RatesLogistic4:
RatesLogistic4 <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)),{
# Optional: forcing functions: get value of the driving variables at time t (if any)
# see template 2
# Optional: auxiliary equations
# Rate equations
dNdt <- r*N*(1-N/K)
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(dNdt = dNdt)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
dINdt <- r*N
dOUTdt <- r*N*N/K
MB <- c(dINdt = dINdt, dOUTdt = dOUTdt)
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
# here we pass the rate of change of N with respect to time back as auxiliary variable
AUX <- RATES
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(c(RATES, # the rates of change of the state variables
MB), # the rates of change of the mass balance terms if MB is not NULL
AUX) # optional additional output per time step
return(outList)
}
)
}Solving the model
Because we will need the initial conditions both while solving the
model, as well as when computing the mass balance checks, let’s store
the initial conditions in an object called inits:
inits <- InitLogistic4()Now we can solve the model, passing inits as input to
argument y:
states <- ode(y = inits,
times = seq(from = 0, to = 50, by = 1),
func = RatesLogistic4,
parms = ParmsLogistic4(),
method = "ode45")Mass balances
To check how the mass balance varies over time, we first need to
specify a function that computes the mass balance as function of the
solved model (states as computed above), as well as the
initial state condition (object inits) as specified above.
In this new function, let’s call it MassBalanceLogistic4,
the solved model will be assigned to input argument y, and
the initial state will be assigned to input argument ini.
Within the function (also here using the with function to
make it easier to access all initial states and columns in our solved
model output directly by their name), we compute the mass balance (thus
in this case the value of at any
moment in time, minus plus the
sum of outflows minus the sum of inflows):
MassBalanceLogistic4 <- function(y, ini) {
# Add "0" to the state variables' names to indicate initial condition
names(ini) = paste0(names(ini), "0")
# Using the with() function we can access elements directly by there name
# here: 'y' is a deSolve matrix
# so it is best to first convert it to a data.frame using function as.data.frame()
# and then combine it with 'ini' using the function c()
# and convert to a list using as.list()
with(as.list(c(as.data.frame(y), ini)), {
# Compute mass balance terms
BAL <- N - N0 + OUT - IN
# Return output BAL
return(BAL)
})
}After specifying this function, we can get the mass balance as a time series and plot to check it:
bal <- MassBalanceLogistic4(y = states, ini = inits)
plot(states[,"time"], bal, type = "l", xlab = "Time", ylab = "Mass balance")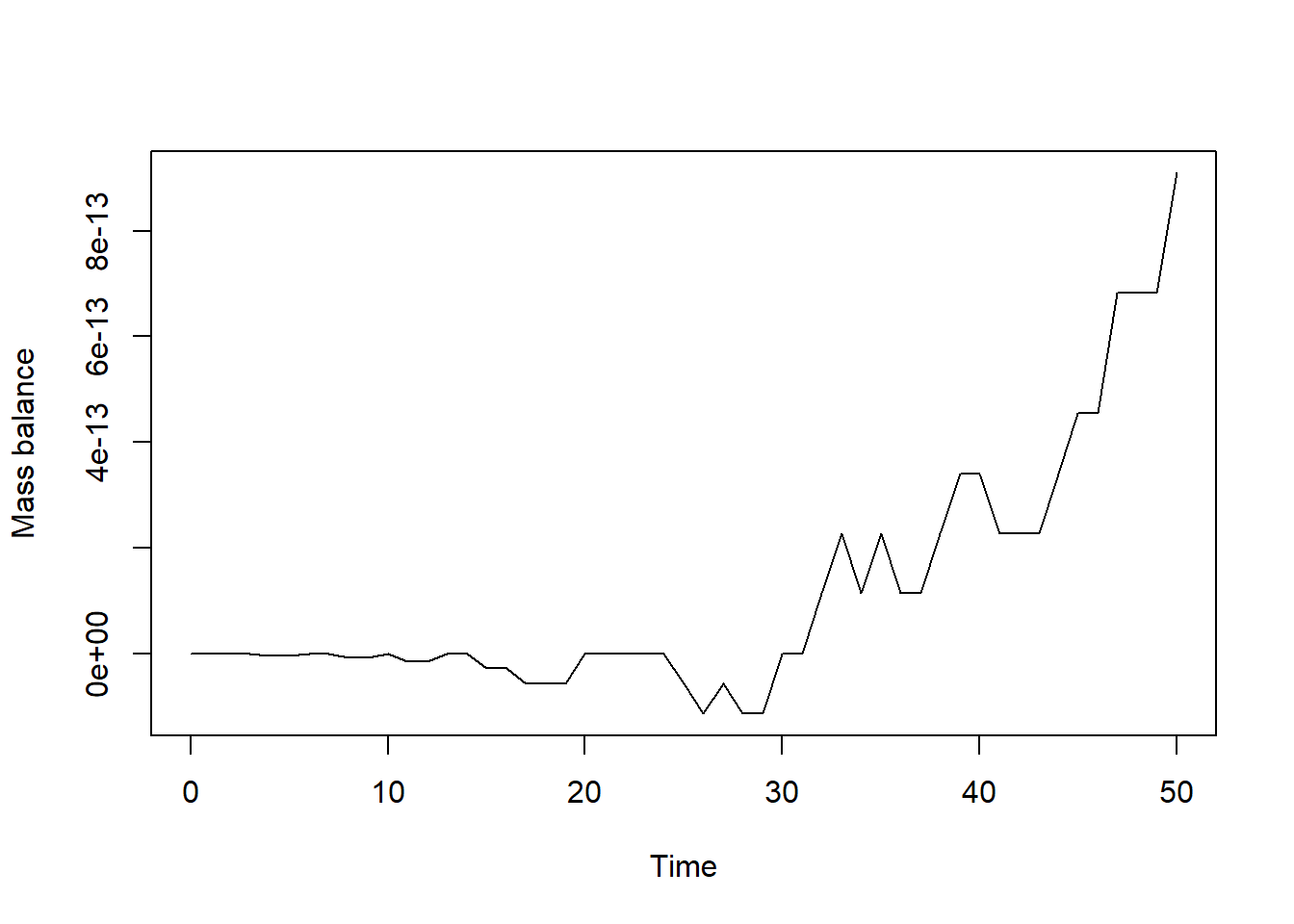
In the plot we see how the mass balance fluctuates over time. It is
sufficiently small (1e-12 equals 0.000000000001!) and not
exactly 0 due to the numerical approximation (and due to rounding error
and the precision with which computers store data). Apart from this, the
mass balance thus seems correct for this model!
Downloadable R script
An R script with the code chunks of the above example can be downloaded here.