Wageningen University & Research | FEM-31806 | Models for Ecological Systems | FEM | PPS | WEC
Appendix B - Mass balances with events
In Template 4, we have demonstrated how you can perform mass balance checks.
Template 3 shows a simple method via which you can specify discrete events that act on state variables at specified times. Appendix A details a more versatile way of specifying events, which can either be occurring at specified times, or which can occur when some condition is met. Here, we show how mass balances (see Template 4) can be kept track of in the presence of discrete events acting upon state variables.
Logistic growth
Let’s use the example of logistic growth as defined in Template 1, and include the objects that allow us to do mass balance checks (Template 4):
# Load the deSolve library
library(deSolve)
# Define a function that returns a named vector with initial state values
InitLogistic <- function(N = 5) {
# Gather function arguments in a named vector
y <- c(N = N,
IN = 0, # sum of inputs
OUT = 0) # sum of outputs
# Return the named vector
return(y)
}
# Define a function that returns a named vector with parameter values
ParmsLogistic <- function(r = 0.3, # intrinsic population growth rate
K = 100 # carrying capacity
){
# Gather function arguments in a named vector
y <- c(r = r,
K = K)
# Return the named vector
return(y)
}
# Define a function for the rates of change of the state variables with respect to time
RatesLogistic <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)), {
# Optional: forcing functions: get value of the driving variables at time t (if any)
# see template 2
# Optional: auxiliary equations
# Rate equations
dNdt <- r*N*(1-N/K)
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(dNdt = dNdt)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
dINdt <- r*N
dOUTdt <- r*N*N/K
MB <- c(dINdt = dINdt, dOUTdt = dOUTdt)
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- NULL
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(c(RATES, # the rates of change of the state variables
MB), # the rates of change of the mass balance terms if MB is not NULL
AUX) # optional additional output per time step
return(outList)
}
)
}
# Function to keep track of mass balances
MassBalanceLogistic <- function(y, ini){
# Add "0" to the state variables' names to indicate initial condition
names(ini) = paste0(names(ini), "0")
# Using the with() function we can access elements directly by there name
# here: 'y' is a deSolve matrix
# so it is best to first convert it to a data.frame using function as.data.frame()
# and then combine it with 'ini' using the function c()
# and convert to a list using as.list()
with(as.list(c(as.data.frame(y), ini)), {
# Compute mass balance terms
BAL <- N - N0 + OUT - IN
# Return output BAL
return(BAL)
})
}Mass balance checks
After defining these functions, we can solve the model, here without events, and perform a mass balance check:
# solve the model
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "ode45")
# mass balance check
bal <- MassBalanceLogistic(states, ini = InitLogistic())
plot(states[,"time"], bal, type = "l", xlab = "Time", ylab = "Mass balance")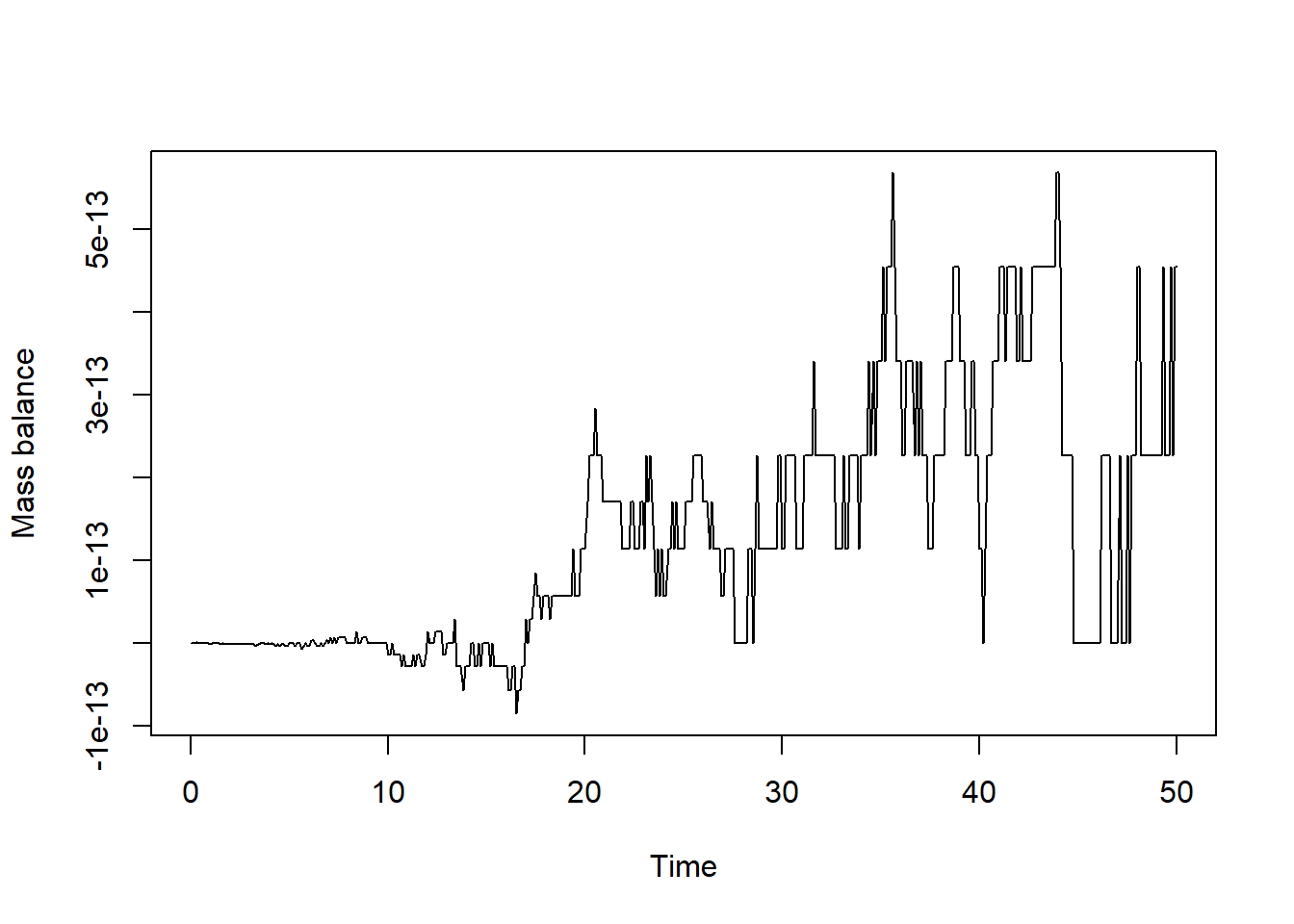
We see that the mass balance is (except for numerical inaccuracy) 0, so this is good!
Harvesting
As in Appendix A, let’s impose some harvesting of this population at pre-defined times: let’s remove 25% of the population at times 10, 20, 30 and 40: we do this via the specification of an event function that is applied at specified times:
# specify event function
eventfun <- function(t, y, parms){
y["N"] <- 0.75 * y["N"]
return(y)
}
# solve the model
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "ode45",
events = list(func = eventfun, # the event function, called at times in "time"
time = c(10,20,30,40), # the time at which the event function is applied
root = FALSE)) # this specifies that we do not supply a root function
# thus: the event function is called at the times supplied in "time"
# plot the simulation
plot(states)
# mass balance check
bal <- MassBalanceLogistic(states, ini = InitLogistic())
plot(states[,"time"], bal, type = "l", xlab = "Time", ylab = "Mass balance")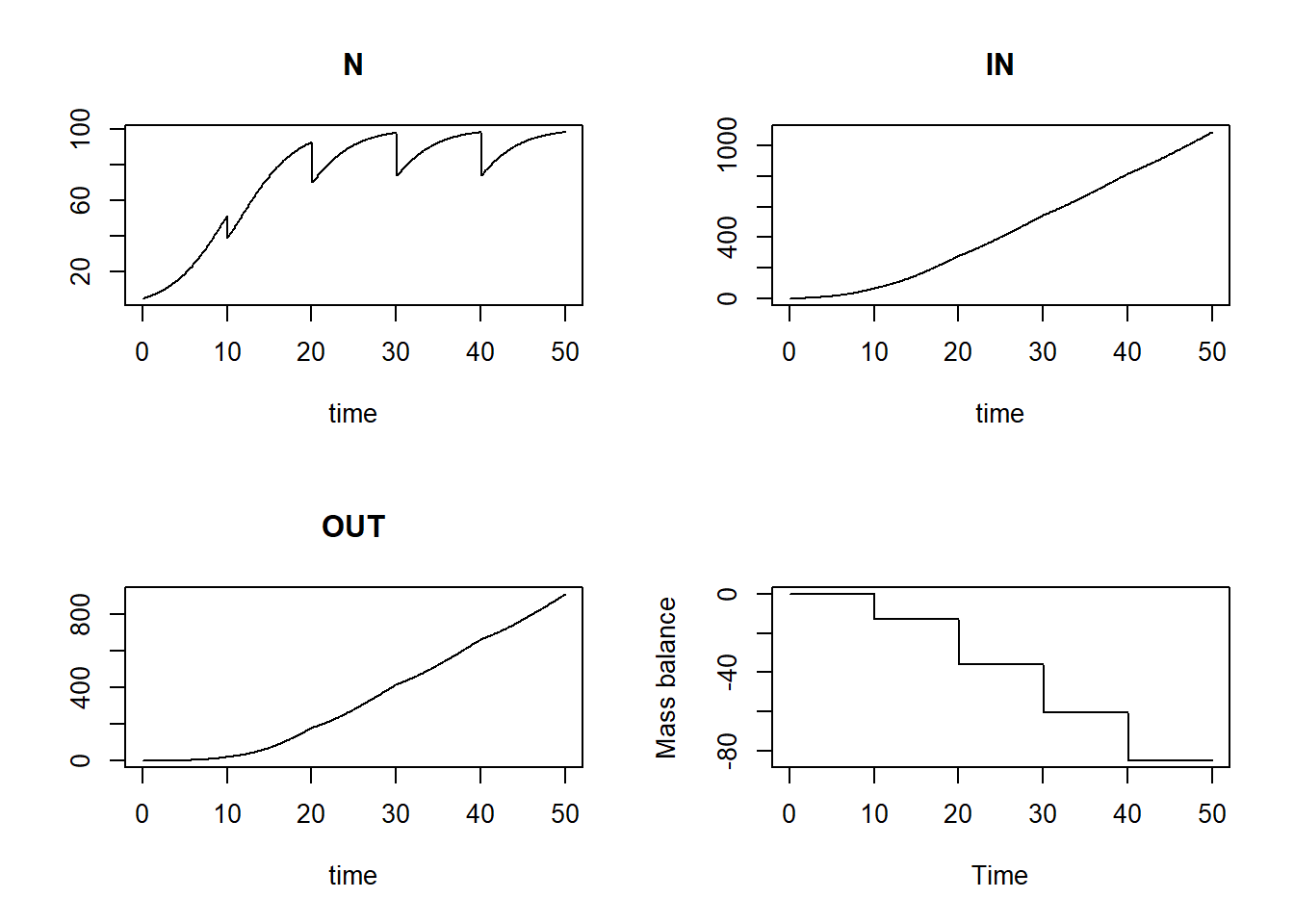
We see that the mass balance is not 0 anymore: we have partly harvested the population at specified times, but this is not considered in the mass balance check, therefore the mass balance is now wrong as it is not 0 anymore!
Mass balances when harvesting
In order to consider the discrete events acting on state variables in
the mass balance check, we need to update the event function so
that it also includes the influence of the event on the mass balance
variables IN and OUT (depending on the type of
event) as defined in the function InitLogistic. As we are
dealing here with reduction of the state due to harvesting, we have to
include the harvested part of the population in the rate of change for
OUT.
Be very careful in the order in which you apply changes to the vector with state variables in the event function! First update the mass balance terms and only then update the state variables.
If done in the wrong order, the mass balance may not be correct
anymore! For example, if we first were to remove 25% from population
at pre-defined times, and only then
add 25% of to the mass balance term
OUT, we actually only add 25% of the population after
reducing it with 25%!
Thus, in the event function eventfun, we first update
the mass balance term OUT (i.e., we add 25% of state to the current value of OUT),
after which we update the value of state (we remove 25% by multiplying it with
0.75):
# specify event function
eventfun <- function(t, y, parms){
# first update OUT: add 25% of N to OUT
y["OUT"] <- y["OUT"] + 0.25 * y["N"]
# only then update N by removing 25%
y["N"] <- 0.75 * y["N"]
# return the updated version of vector y
return(y)
}
# solve the model
states <- ode(y = InitLogistic(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesLogistic,
parms = ParmsLogistic(),
method = "ode45",
events = list(func = eventfun, # the event function, called at times in "time"
time = c(10,20,30,40), # the time at which the event function is applied
root = FALSE)) # this specifies that we do supply a root function
# plot the simulation
plot(states)
# mass balance check
bal <- MassBalanceLogistic(states, ini = InitLogistic())
plot(states[,"time"], bal, type = "l", xlab = "Time", ylab = "Mass balance")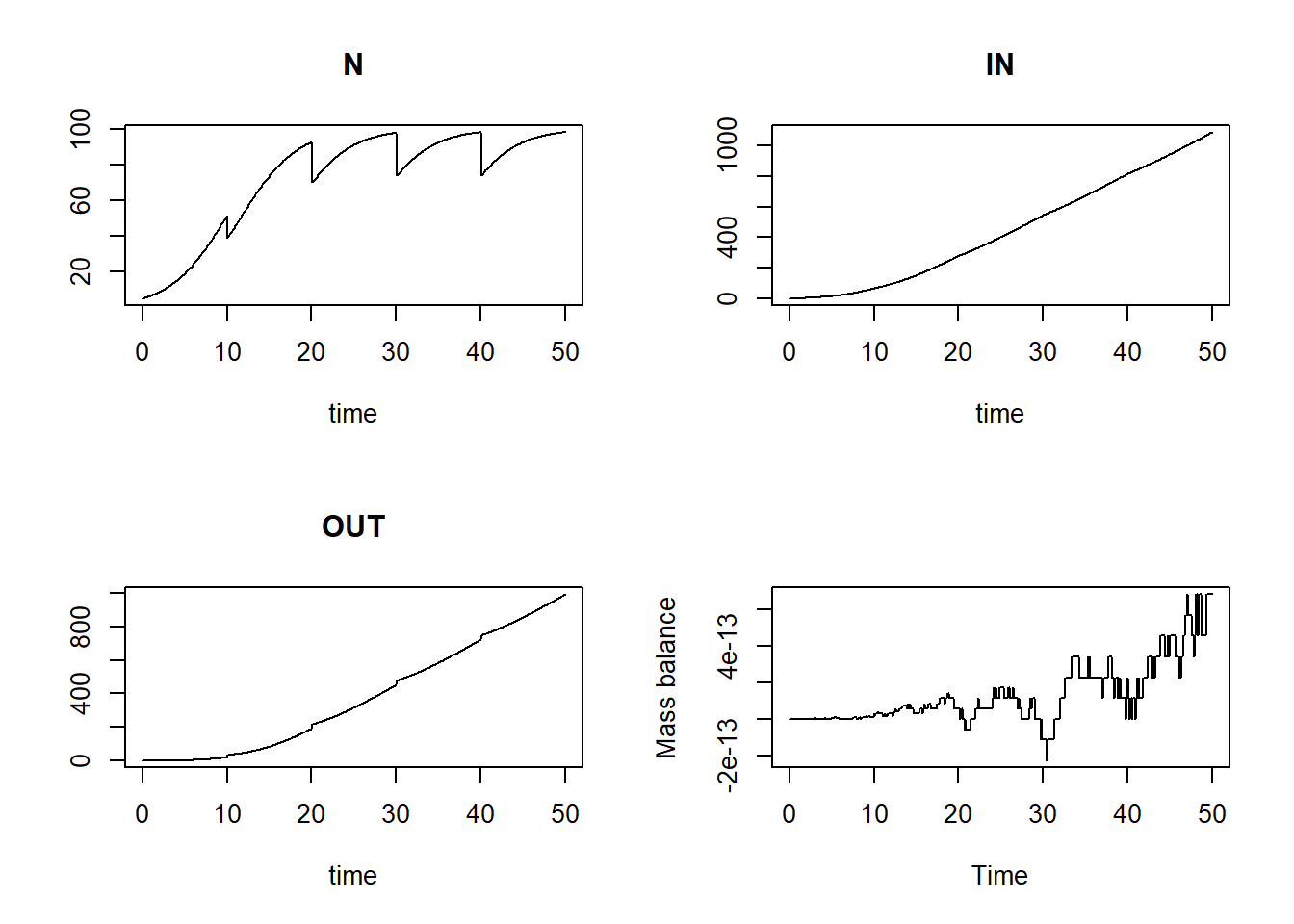
Now we see that the mass balance is again ca. 0 across the entire simulation, as it should!
Downloadable R script
An R script with the code chunks of the above example can be downloaded here.