Wageningen University & Research | FEM-31806 | Models for Ecological Systems | FEM | PPS | WEC
Chapter 4 - Numerical integration
Knowledge questions
Numerical integration is preferred when an analytical (closed‑form) solution is impossible, impractical, or inefficient to obtain.
This typically happens in situations such as:
- The integrand is too complex so that an analytical solution is not possible (or even when a closed-form solution might exist in theory, it may be extremely complicated or computationally expensive to derive);
- The integrand has irratic behaviour, such as sharp peaks, discontinuities (e.g. in the case of discrete event), or highly oscillatory behaviour. This makes analytical integration difficult, while numerical methods can adapt to these features;
- The system includes a driving variable: external inputs (e.g. rainfall) often appear as arbitrary functions of time, and often are non-smooth and irregular, so that analytical integration becomes impossible.
Exercises
Have a look at the Templates section of this website, where we provide templates to solve ODE models using R. Following the naming convention used in the Templates, let’s call the function in this exercise “PredatorPrey”, thus below you will be defining the functions needed to conveniently solve the models and give them names like “InitPredatorPrey”, “ParmsPredatorPrey”, and “RatesPredatorPrey”.
In the syllabus and lecture, we were modelling the dynamics of a single population, but now we will be modelling a predator-prey system. The Lotka-Volterra model is a cornerstone for the theory about predator-prey interactions, and dates back to the 1920s. The basic Lotka-Volterra predator-prey model is based on the following assumptions:
- prey would multiply indefinitely (i.e. grow exponentially) in the absence of predators;
- predators would go extinct if prey were absent, due to a lack of food;
- the proportional rate of increase of the prey decreases as the number of predators increases;
- the growth rate of predators increases when the number of prey increases.
Using these assumptions, we can model the predator-prey system with a simple set of ordinary differential equations (ODE’s):
where and represent the abundance of prey (Resource) and predators (Consumers), respectively. The parameter represents the exponential growth rate of prey in the absence of the predator, while represents the mortality rate of the predators in the absence of prey. Encounters between prey and predators are assumed to follow the mass action law and are hence proportional to the product of the abundances and . The parameter represents the attack/kill rate. The parameter represents the conversion efficiency, i.e. the efficiency with which predators convert consumed prey into biomass.
Exercise 4.1
Start RStudio and open a new script. Save the script at some
convenient place. Load the package deSolve (install the
package when it is not installed yet). In this exercise you are going to
implement the predator-prey model as described above. Therefore, use the
same names for the state variables (
and ) and the parameters (, , , ), and
call the rate variables that you will compute and . Information (and examples) that can be
helpful are found in section 4.3 of the syllabus, as well as the
template files provided on this website to solve ODE models numerically
(for this exercise you can use
Template 1).
InitPredatorPrey,
and specify the default values 0.5 for prey (or resource: ), and 0.1 for predator (or consumer: ). InitPredatorPrey <- function(R = 0.5, # Resource / prey
C = 0.1){ # Consumer / predator
# Gather function arguments in a named vector
y <- c(
R = R,
C = C
)
# Return the named vector
return(y)
}ParmsPredatorPrey, and set as default values: 0.5 for , 1.0 for , 0.5 for
and 0.1 for . ParmsPredatorPrey <- function(g = 0.5, # exponential growth rate of prey in the absence of the predator
b = 1.0, # attack/kill rate
c = 0.5, # conversion efficiency
m = 0.1){ # mortality rate of the predators in the absence of prey
# Gather function arguments in a named vector
y <- c(
g = g,
b = b,
c = c,
m = m
)
# Return the named vector
return(y)
}RatesPredatorPrey, that
computes the rates of change of the state parameters, and that will be
evaluated by the ode() function. Remember that the
ode() function requires the function to have as function
arguments: t, y, parms (in that
order, but you can use your own argument names, e.g. Time, State and
Parms). Moreover, the function should return a list with as
first element the rates of change of the state variables (here there are
2, so combine them in a vector using the function c()), in
the order as specified in InitPredatorPrey. RatesPredatorPrey <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# thus: 'y' and 'parms' are combined using function c and converted using function as.list
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)), {
# Optional: forcing functions: get value of the driving variables at time t (if any)
# Optional: auxiliary equations
# Rate equations
dRdt <- g * R - b * R * C
dCdt <- c * b * R * C - m * C
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(
dRdt = dRdt,
dCdt = dCdt
)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- NULL
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(
c(
RATES, # the rates of change of the state variables
MB
), # the rates of change of the mass balance terms if MB is not NULL
AUX
) # optional additional output per time step
return(outList)
}
)
}tseq. Use the function seq() for this.
tseq <- seq(from = 0, to = 100, by = 0.5)ode() in the deSolve package to
numerically solve the predator-prey model. Use the initial conditions
(the function InitPredatorPrey created in (a)), parameter
values (the function ParmsPredatorPrey as created in (b))
and function RatesPredatorPrey as created in (c). Use the
variable tseq as input to the times argument in
the call to the function ode(). Assign the solved ODE to
the object states. Use “ode45” as the integration method
supplied to method, i.e. an adaptive stepsize Runge-Kutta
integration method (also in the following exercises when solving a model
numerically). states <- ode(
y = InitPredatorPrey(),
times = tseq,
func = RatesPredatorPrey,
parms = ParmsPredatorPrey(),
method = "ode45"
)states (which
essentially is a matrix) using the functions head() and
tail(). What are the column names, and how do they relate
to the initial state conditions as specified in
InitPredatorPrey as created in exercise (a)? head(states)## time R C
## [1,] 0.0 0.5000000 0.1000000
## [2,] 0.5 0.6093871 0.1092289
## [3,] 1.0 0.7385141 0.1229174
## [4,] 1.5 0.8875292 0.1432181
## [5,] 2.0 1.0533754 0.1735900
## [6,] 2.5 1.2269235 0.2195823tail(states)## time R C
## [196,] 97.5 0.0014499409 0.8123317
## [197,] 98.0 0.0012526686 0.7729740
## [198,] 98.5 0.0011032233 0.7354916
## [199,] 99.0 0.0009895405 0.6998038
## [200,] 99.5 0.0009031619 0.6658312
## [201,] 100.0 0.0008380994 0.6334958We can see that the columns names of the deSolve matrix
states are “time” as well as “R” and “C”: the names of the
states as specified in the input to argument y (the output
of the function call InitPredatorPrey).
states. Simply use
plot(states) to use the default plot method for objects
returned by the function ode. plot(states)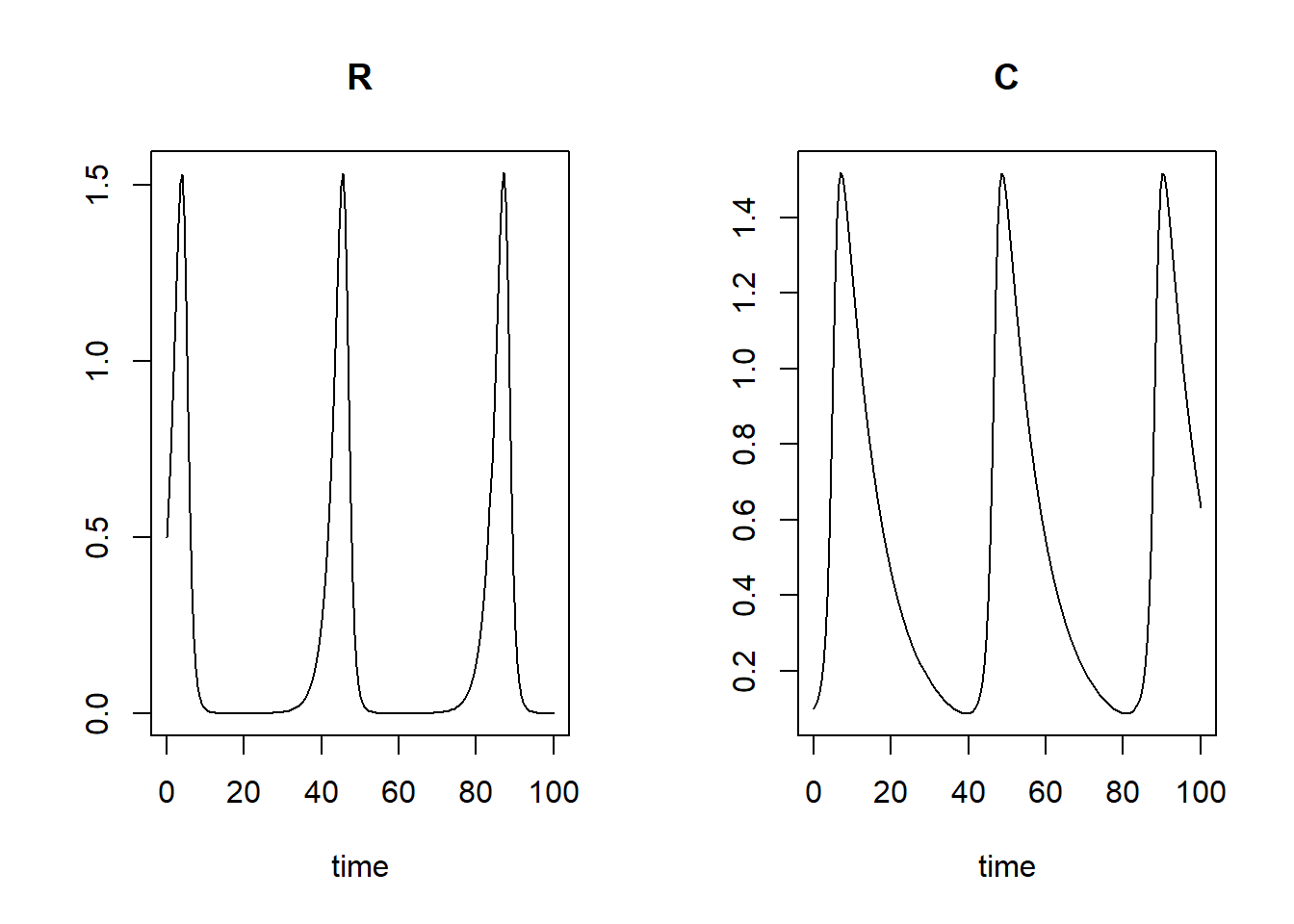
ode() is a
matrix. Also note that you’ve seen in (f) that the column names are
“time”, “R” and “C” (the latter 2 because the function InitPredatorPrey
specifies the 2 initial states to be “R” and “C”!). Remember that the
column “R” can be retrieved using matrix indexing in different ways,
here using either states[,2] or states[,"R"],
because is the second column, and has
the column name “R”. A plot of x-values against y-values can be
generated using plot(x = xvals, y = yvals, type = "l"),
where xvals and yvals are hypothetical objects with
data, and type = "l" specifies that the type of plot that
is generated is a line plot. You can choose to adjust the axis labels by
providing input to the arguments xlab and
ylab. plot(states[, "R"], states[, "C"], type = "l", xlab = "R", ylab = "C")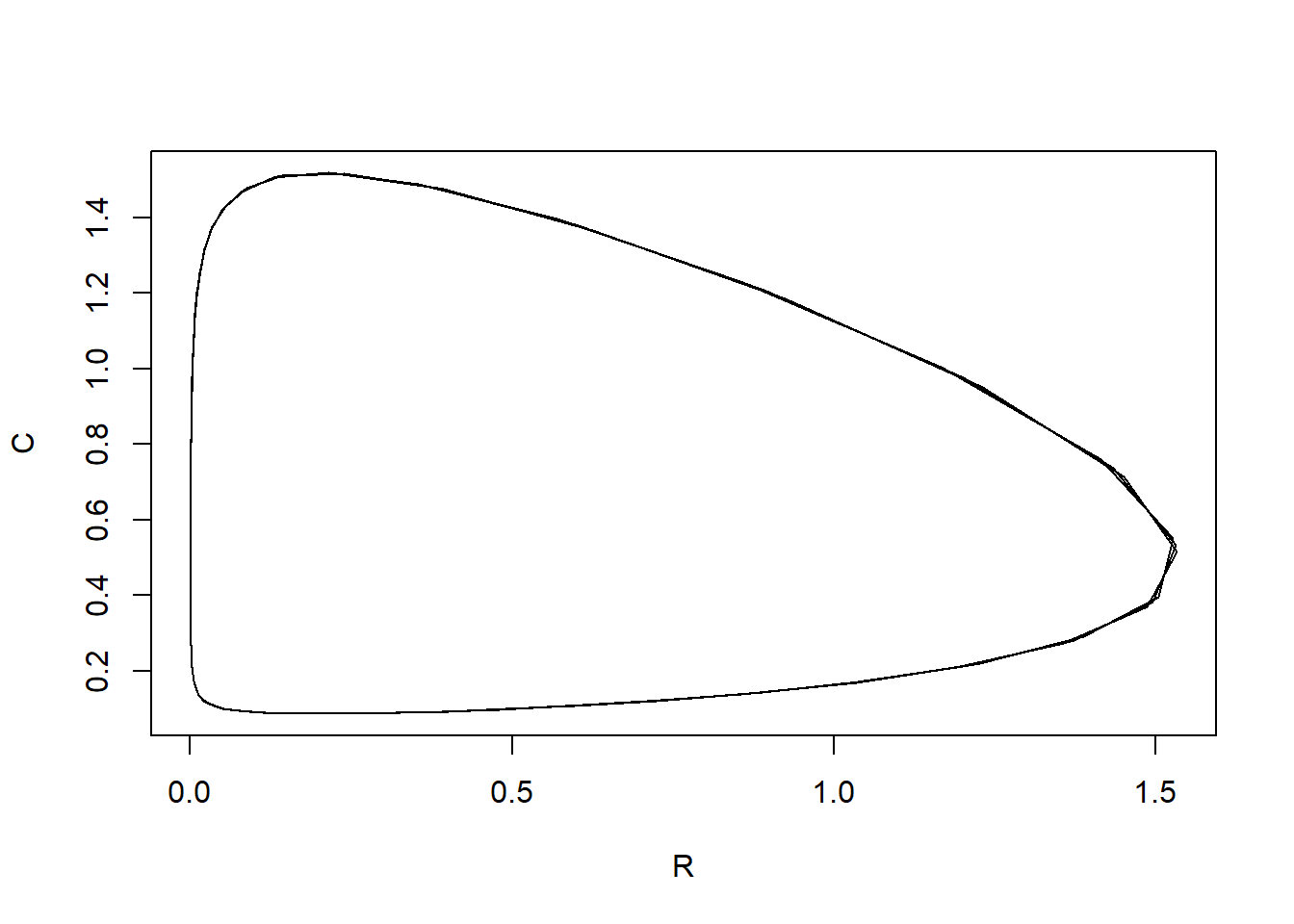
We see that when the resource increases, then also the consumer will increase, which will then cause a decrease in the resource, which will also result in a decrease in the consumer. This is a cycle that repeats itself over and over (the predator-prey cycle).
Note that the plotted line is not perfectly smooth: since we solve
the model with time-steps of 0.5 (see how tseq was
specified above); if we would solve the model with smaller a time-step,
the line should look more smooth).
The nullclines (also called zero-growth isoclines) of the
Lotka-Volterra predator-prey model are known: the nullcline is given and . The nullcline is given by and . Obviously the nullclines
where the states are 0 are not of interest (as we would not have a
predator-prey system anymore). Thus, add the other 2 nullclines to the
plot ( and ). To do this, use the
abline() function, where a line can be drawn horizontally
or vertically at some value by specifying h=value or
v=value, respectively. Alternatively, a linear line that is
not parallel to the x or y axis can be drawn using the input arguments
a=value and b=value, for intercept
a and slope b. Use the same parameter values
as passed to the function ode (thus retrieve these values
from the ParmsPredatorPrey function).
Tip: it may be handy to use of the with function, and
inside this the abline function:
with(as.list(ParmsPredatorPrey()), {
abline(h = g / b, col = 8, lty = 2)
abline(v = m / (b * c), col = 8, lty = 2)
})plot(states[, "R"], states[, "C"], type = "l", xlab = "R", ylab = "C")
with(as.list(ParmsPredatorPrey()), {
abline(h = g / b, col = 8, lty = 2)
abline(v = m / (b * c), col = 8, lty = 2)
})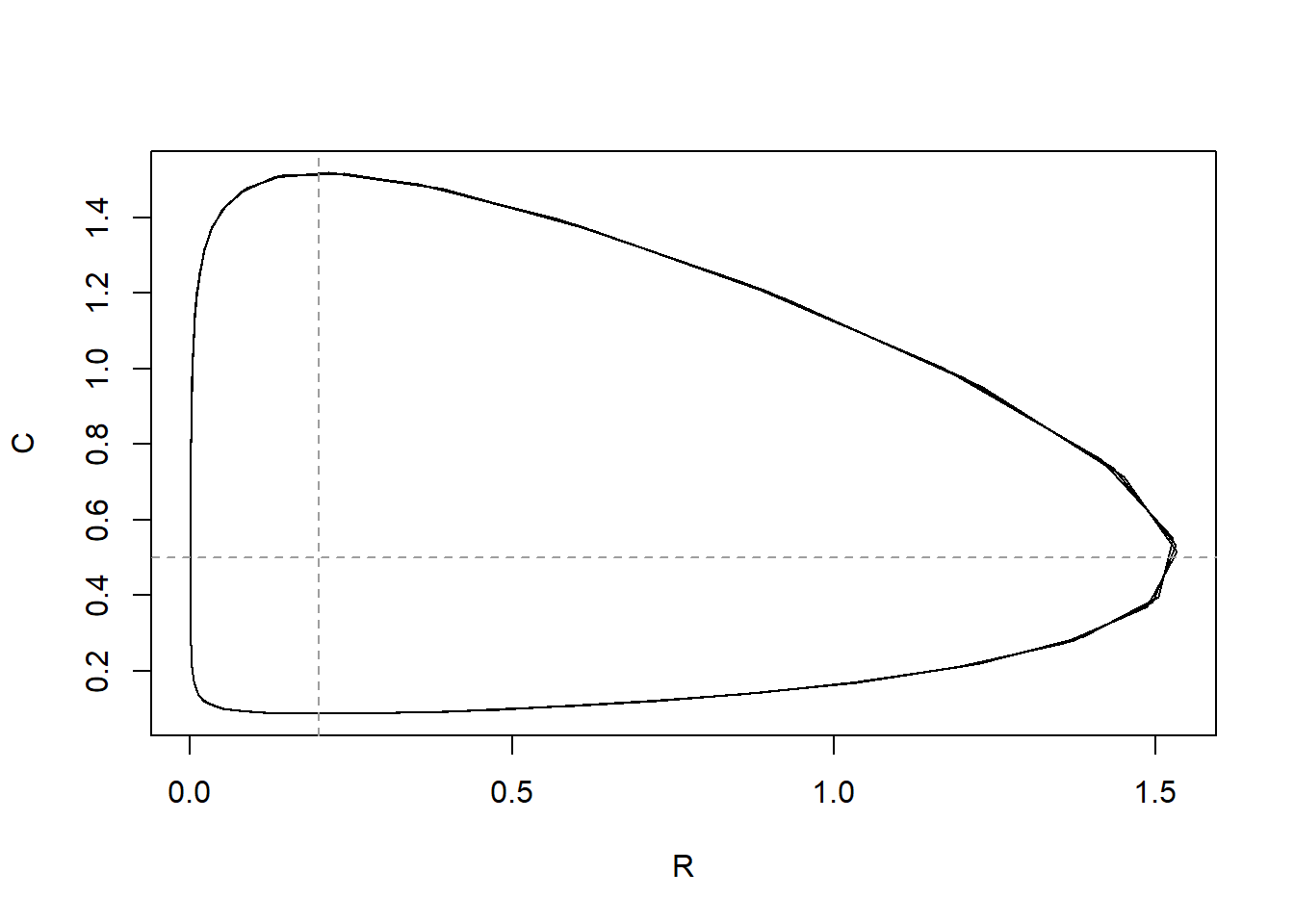
The nullclines indicate the situation where either the resource or the consumer does not change.
Solve the model for a longer duration: to 1000 time units, and assign
it to a new object states2. Add the line of vs to
the plot, using the function lines(). To specify the colour
of the line, use the argument col (which can be a character
string, e.g. “red” or “blue”, but also a value, e.g. 1 is black, 2 is
red, 3 is green, etc). By adding the line in a different colour (than
the current line in the plot), you can identify the added line
easier.
states2 <- ode(
y = InitPredatorPrey(),
times = seq(from = 0, to = 1000, by = 0.5),
func = RatesPredatorPrey,
parms = ParmsPredatorPrey(),
method = "ode45"
)
plot(states[, "R"], states[, "C"], type = "l", xlab = "R", ylab = "C", col = "black")
lines(states2[, "R"], states2[, "C"], col = "red")
with(as.list(ParmsPredatorPrey()), {
abline(h = g / b, col = 8, lty = 2)
abline(v = m / (b * c), col = 8, lty = 2)
})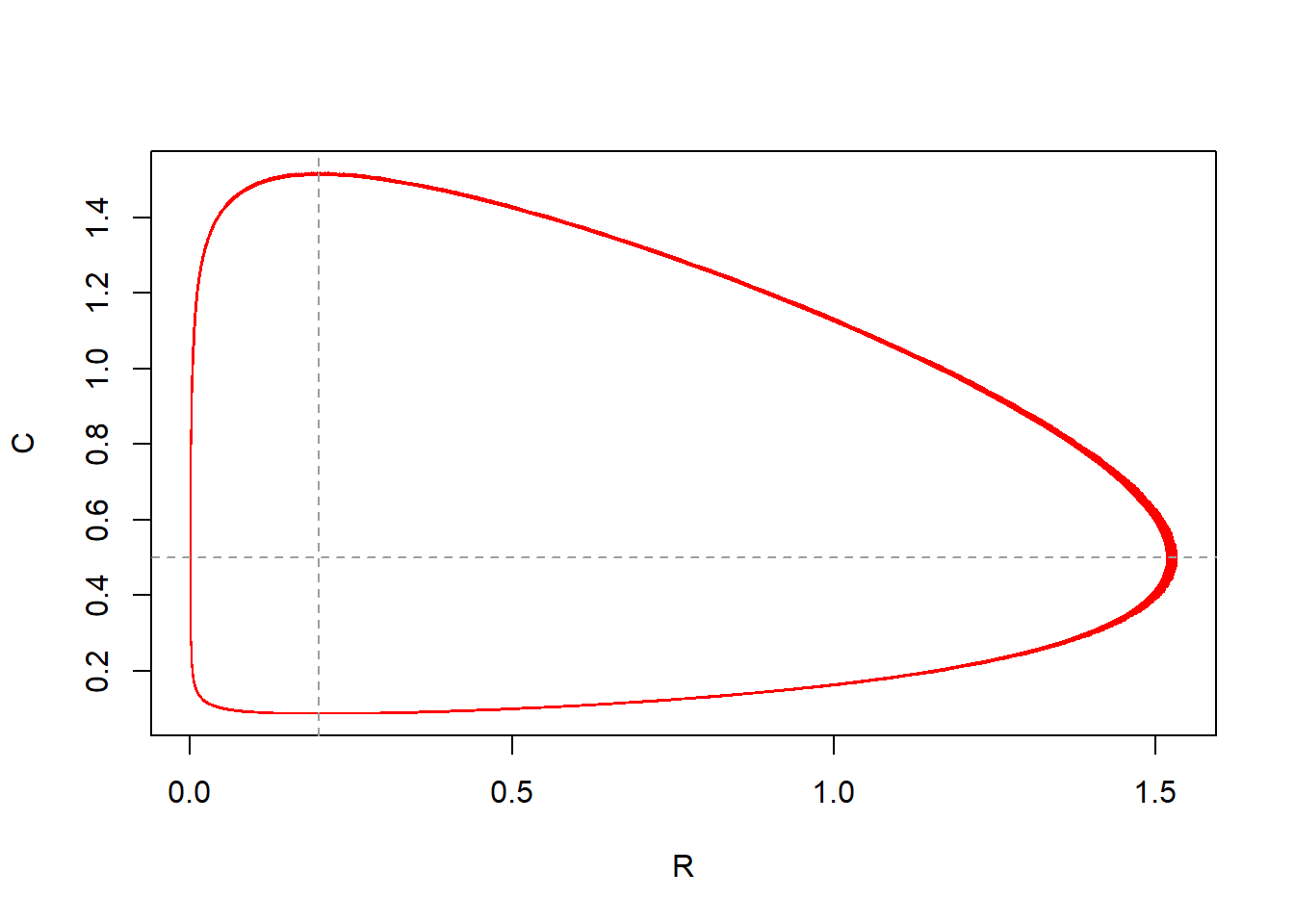
The output of the model solved for 1000 time units nicely fits the pattern as is already visible when solving the model for 100 time units. In the plot the added red line perfectly overlays the red line. Thus: solving the model for longer does not show different patterns; the predator-prey cycle pattern are already captured at 100 time units.
states3), and one
where the initial conditions are ,
(save as states4).
For both scenarios, use the default parameter values and output times as
specified in tseq. Add the lines with different colours so
that you can identify the different scenarios. What can you conclude
from the pattern shown in the plot? How does this match with the
nullclines? If we solve the model for different initial values of and we see that the result changes: the closer we initiate the states to the equilibrium, the smaller the cycles become:
plot(states[, "R"], states[, "C"], type = "l", xlab = "R", ylab = "C")
with(as.list(ParmsPredatorPrey()), {
abline(h = g / b, col = 8, lty = 2)
abline(v = m / (b * c), col = 8, lty = 2)
})
states2 <- ode(
y = InitPredatorPrey(),
times = seq(from = 0, to = 1000, by = 0.5),
func = RatesPredatorPrey,
parms = ParmsPredatorPrey(),
method = "ode45"
)
lines(states2[, "R"], states2[, "C"], col = 2)
states3 <- ode(
y = InitPredatorPrey(R = 0.25, C = 0.4),
times = tseq,
func = RatesPredatorPrey,
parms = ParmsPredatorPrey(),
method = "ode45"
)
lines(states3[, "R"], states3[, "C"], col = 4)
states4 <- ode(
y = InitPredatorPrey(R = 0.2, C = 0.45),
times = tseq,
func = RatesPredatorPrey,
parms = ParmsPredatorPrey(),
method = "ode45"
)
lines(states4[, "R"], states4[, "C"], col = 3)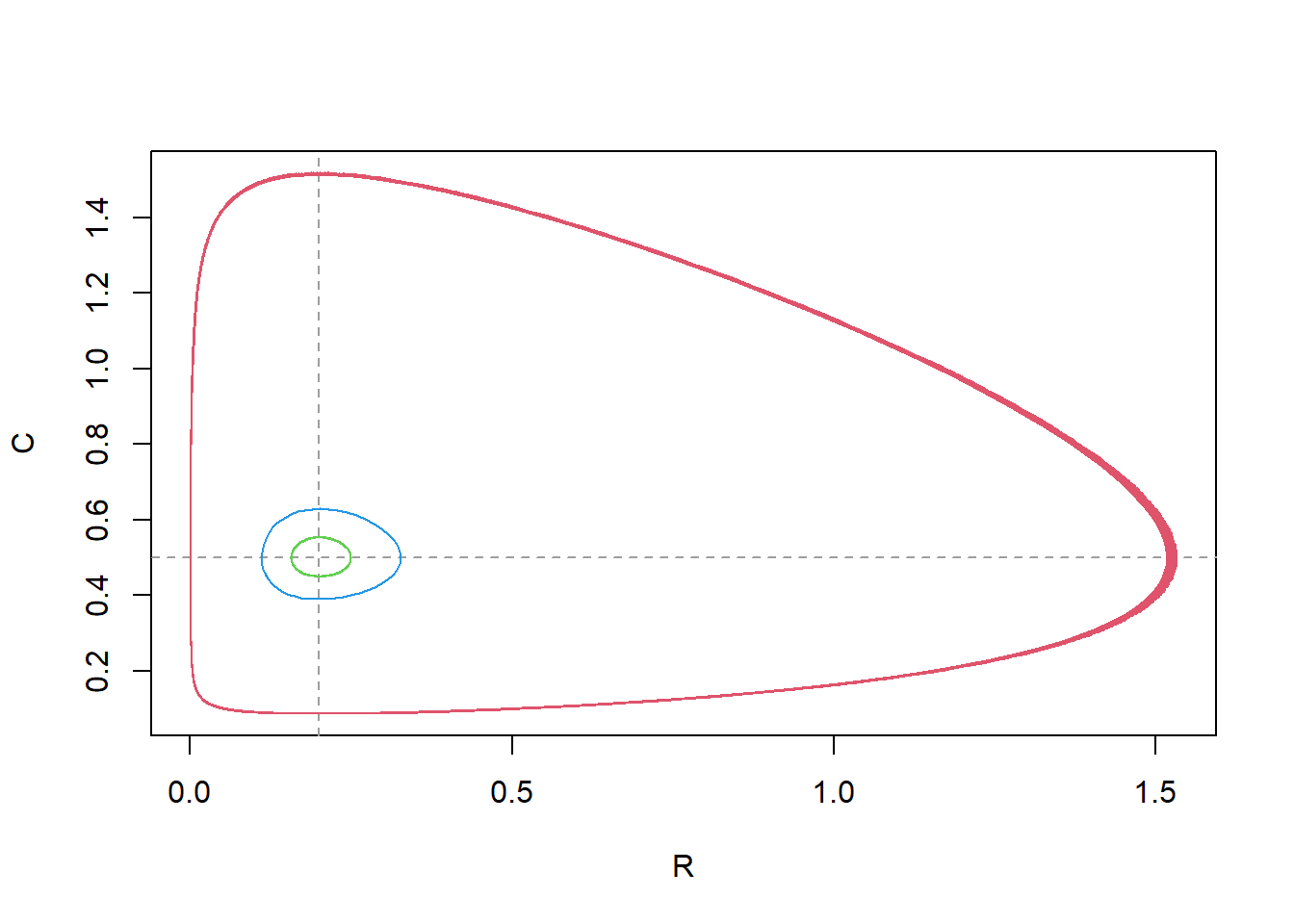
tseq)
and save as states5. Plot the solution. When we first compute the equilibrium values of the states, and then initiate a simulation with those values, we see that the model stays in equilibrium throughout the simulation (what it is supposed to do):
stableInits <- with(as.list(ParmsPredatorPrey()), {
c(R = m / (b * c), C = g / b)
})
states5 <- ode(
y = stableInits,
times = tseq,
func = RatesPredatorPrey,
parms = ParmsPredatorPrey(),
method = "ode45"
)
plot(states5)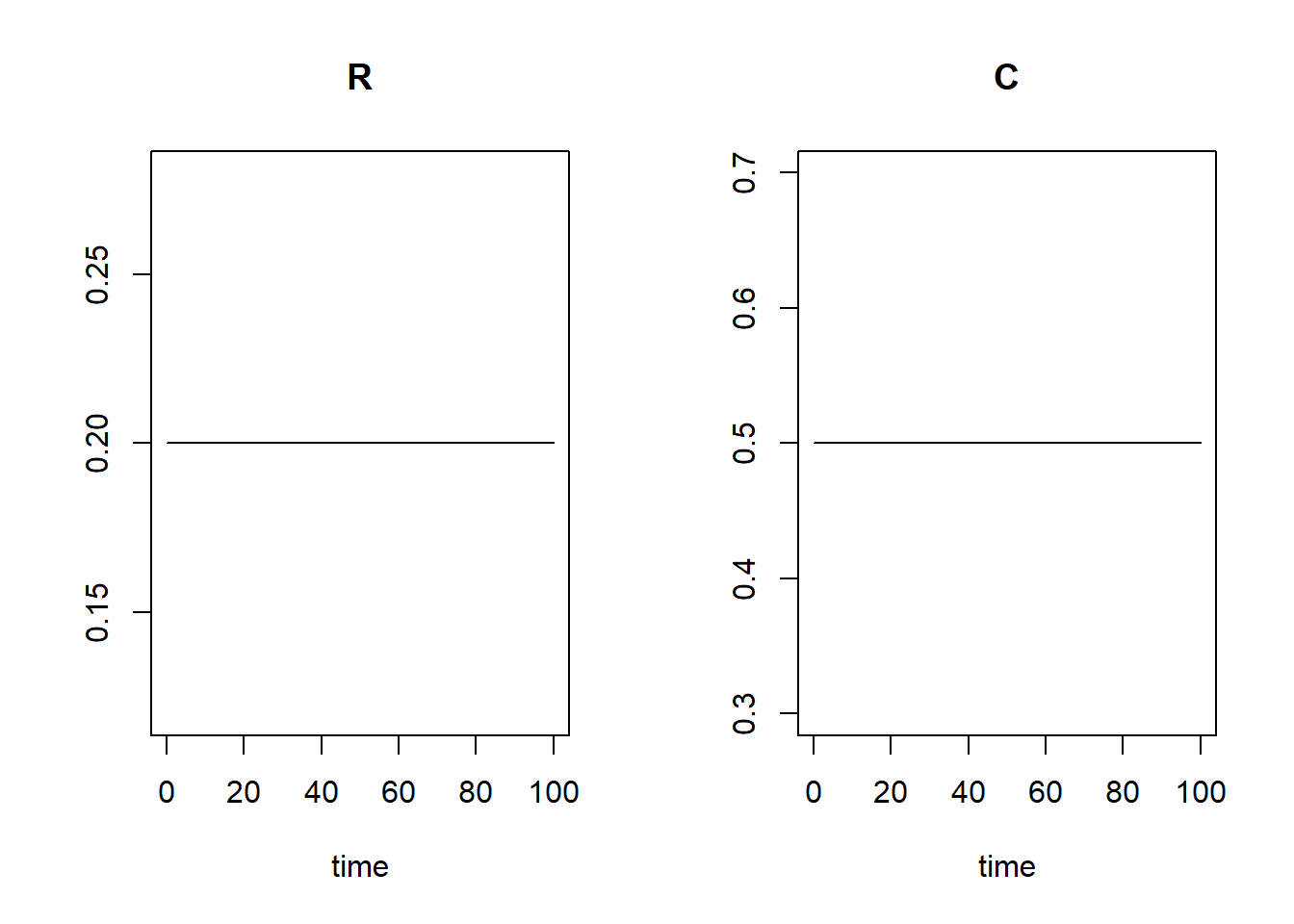
states, states3, states4 and
states5 in a single plot to compare the different scenarios
(all evaluated on the same time scale tseq). Remember that
the deSolve package allows for easy comparison of different scenarios by
allowing multiple objects returned by the ode() function in
a call to the plot() function,
e.g. plot(states,states2). We can plot multiple model simulations in a single plot:
plot(states, states3, states4, states5)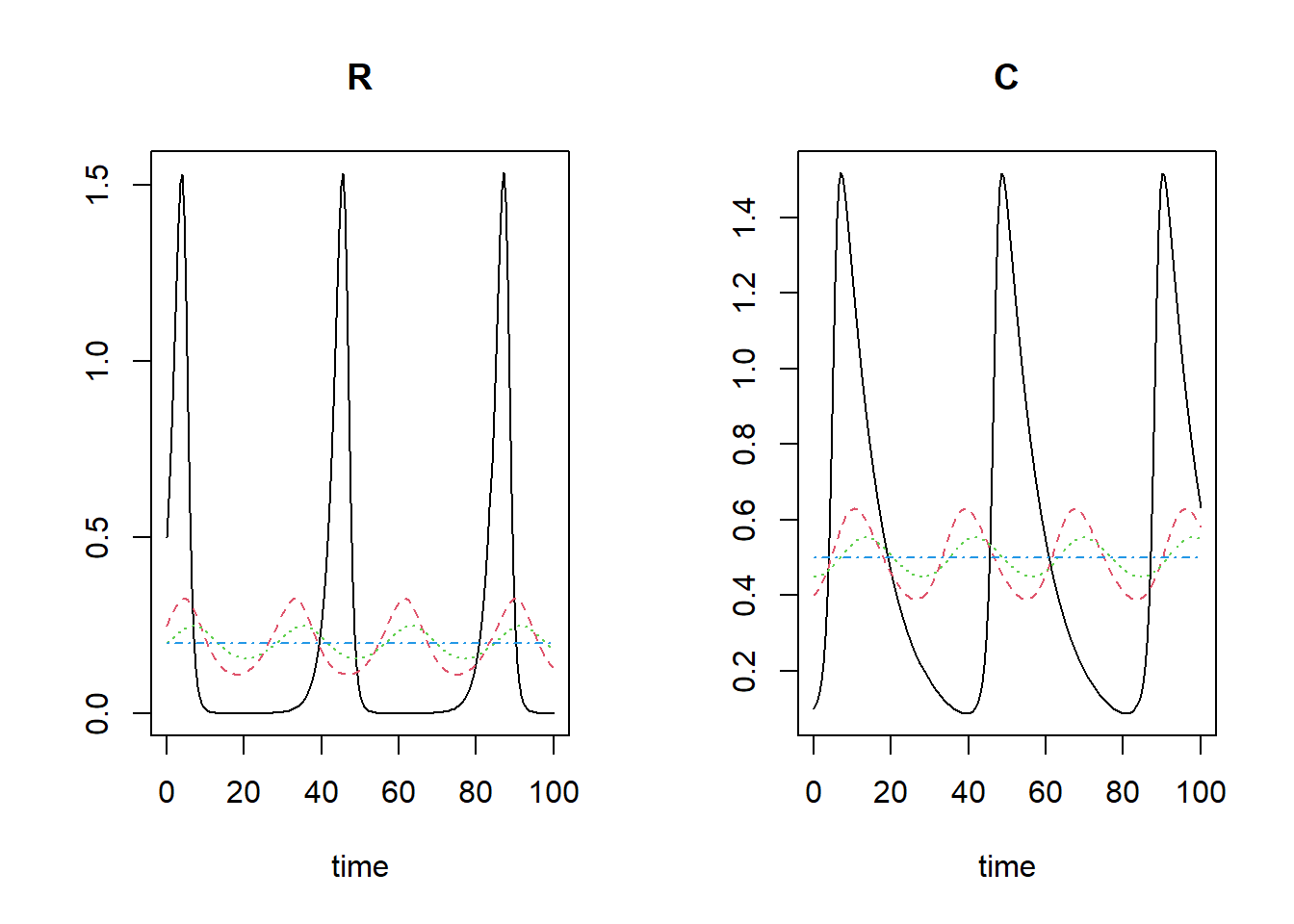
Note that the line colours and line-types follow the standard R colour/line-types 1:n, thus: the black line is for the first model, the red line is for the second model, the green line for model 3, blue for model 4.
Optional exercises
Exercise 4.2
A slightly more complicated and realistic version of the Lotka-Volterra predator-prey model assumes that prey do not grow indefinitely in the absence of predators but will ultimately reach a maximum prey abundance. Hence, prey do not grow exponentially, but follow, for example, a logistic growth equation, like we saw in the populations modelled in the syllabus and lectures. We can thus update the ODEs of the predator-prey model:
The only difference of this set of equations compared to the previous model is that we replaced the parameter with the term , characterizing the logistic growth process of prey up to a carrying capacity .
InitPredatorPrey), whereas other parts
(e.g. the functions ParmsPredatorPrey, and
RatesPredatorPrey) should be updated (use the default
values 0.5 for and 10 for ). Give all functions now a new name,
e.g. InitPredatorPrey2 etc. Solve the predator-prey system
using the ode() function, using time-series stretching 100
time units (starting from 0 and ending at 100), but also longer
(e.g. ending at 1000), and plot the results (both the default plot
method, but also a plot with prey on
x-axis vs. predator on y-axis). What
is the main difference compared to the predator-prey system dynamics as
seen in exercise 4.1? InitPredatorPrey2 <- function(R = 0.5, # Resource / prey
C = 0.1){ # Consumer / predator)
# Gather function arguments in a named vector
y <- c(
R = R,
C = C
)
# Return the named vector
return(y)
}
ParmsPredatorPrey2 <- function(r = 0.5, # intrinsic growth rate of logistic growth
K = 10, # carrying capacity
b = 1.0, # attack/kill rate
c = 0.5, # conversion efficiency
m = 0.1){ # mortality rate of the predators in the absence of prey
# Gather function arguments in a named vector
y <- c(
r = r,
K = K,
b = b,
c = c,
m = m
)
# Return the named vector
return(y)
}
# Differences compared to exercise 4.1:
# - the rate equations
RatesPredatorPrey2 <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# (thus combine 'y' and 'parms' first, using the function c(), then convert to list using as.list())
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)), {
# Optional: forcing functions: get value of the driving variables at time t (if any)
# Optional: auxiliary equations
# Rate equations
dRdt <- r * R * (1 - R / K) - b * R * C
dCdt <- c * b * R * C - m * C
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(
dRdt = dRdt,
dCdt = dCdt
)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- NULL
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(
c(RATES, MB), # the rates of change
AUX
) # additional output per time step
return(outList)
}
)
}
states <- ode(
y = InitPredatorPrey2(),
times = seq(from = 0, to = 1000, by = 0.5),
func = RatesPredatorPrey2,
parms = ParmsPredatorPrey2(),
method = "ode45"
)
plot(states)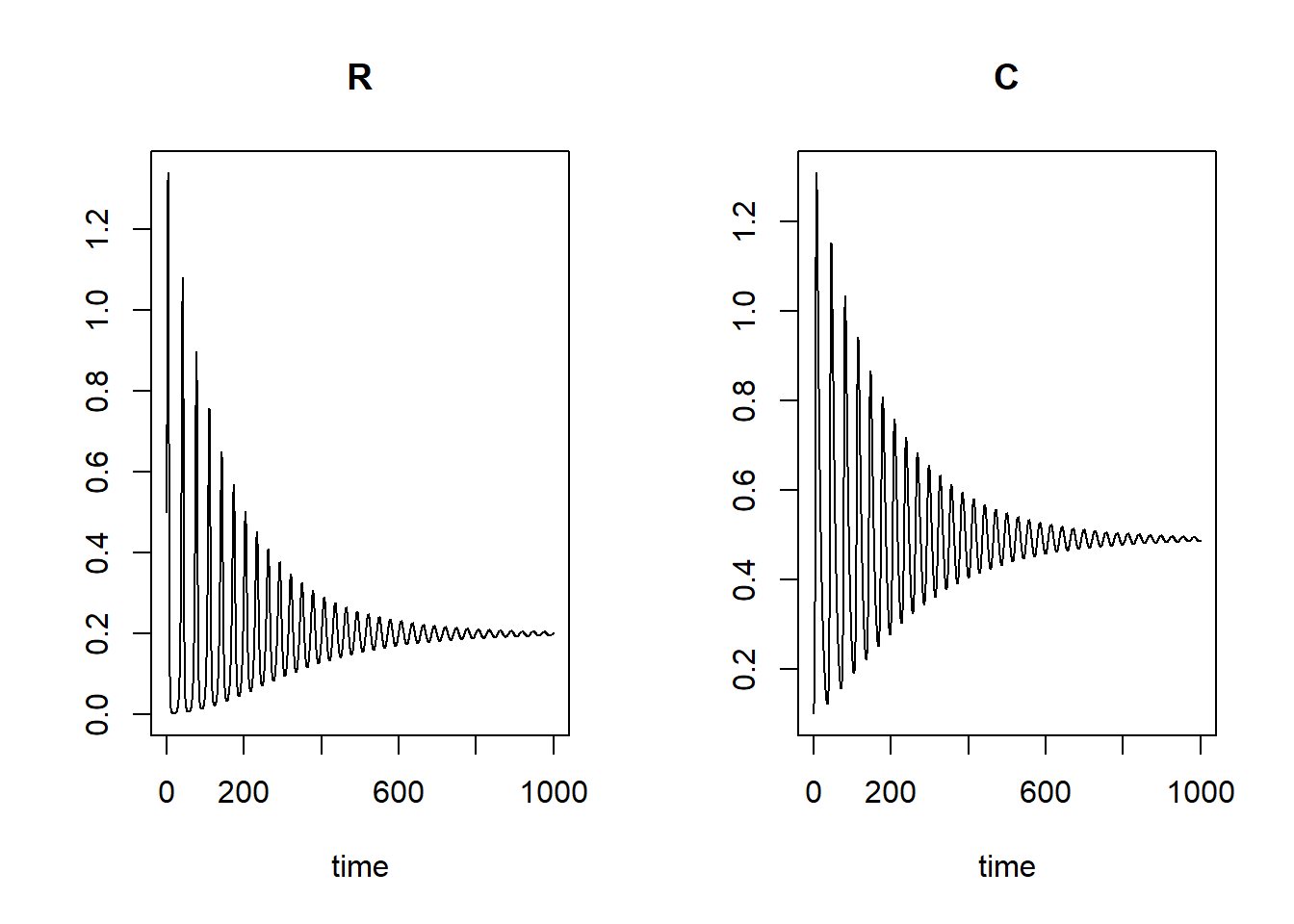
plot(states[, "R"], states[, "C"], type = "l", xlab = "R", ylab = "C")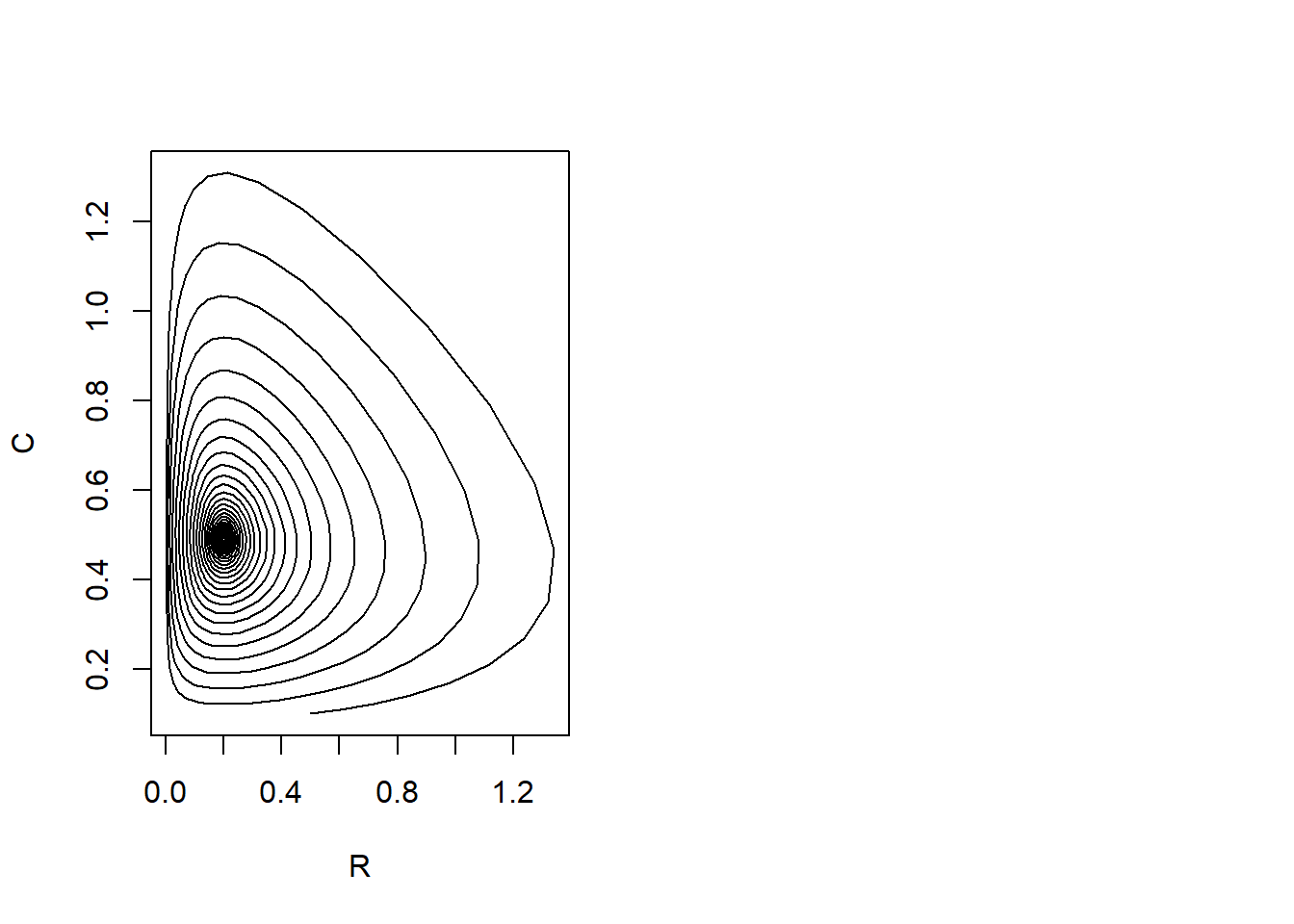
We can see that the main difference of the model from the previous exercises is that in this model the predator and prey will gradually oscillate to an equilibrium value: the cycles we saw in the previou exercises are getting smaller and smaller.
seq function is handy here). Then, create an empty
list called outlist and iterate through the different
scenarios (as specified in the vector kseq) using a
for loop (see section 4.5 of the syllabus for examples, as
well as section 3.11 for information on the use of loops). In each
iteration i of the loop, use the i-th element
of the vector kseq as the value for , solve the ODE, and store the output in the
i-th element of the list outlist (use double
square brackets!). Then, plot the output using the plot method that
deSolve offers for plotting multiple scenarios. Because you store all
the scenarios in a single list, the following code will do the job:
plot(outlist[[1]],outlist[-1]). This will create a plot
with all scenarios. kseq <- seq(from = 0.01, to = 10, length = 10)
outlist <- list()
for (i in 1:length(kseq)) {
outlist[[i]] <- ode(
y = InitPredatorPrey2(),
times = seq(from = 0, to = 1000, by = 1),
func = RatesPredatorPrey2,
parms = ParmsPredatorPrey2(K = kseq[i]),
method = "ode45"
)
}
plot(outlist[[1]], outlist[-1])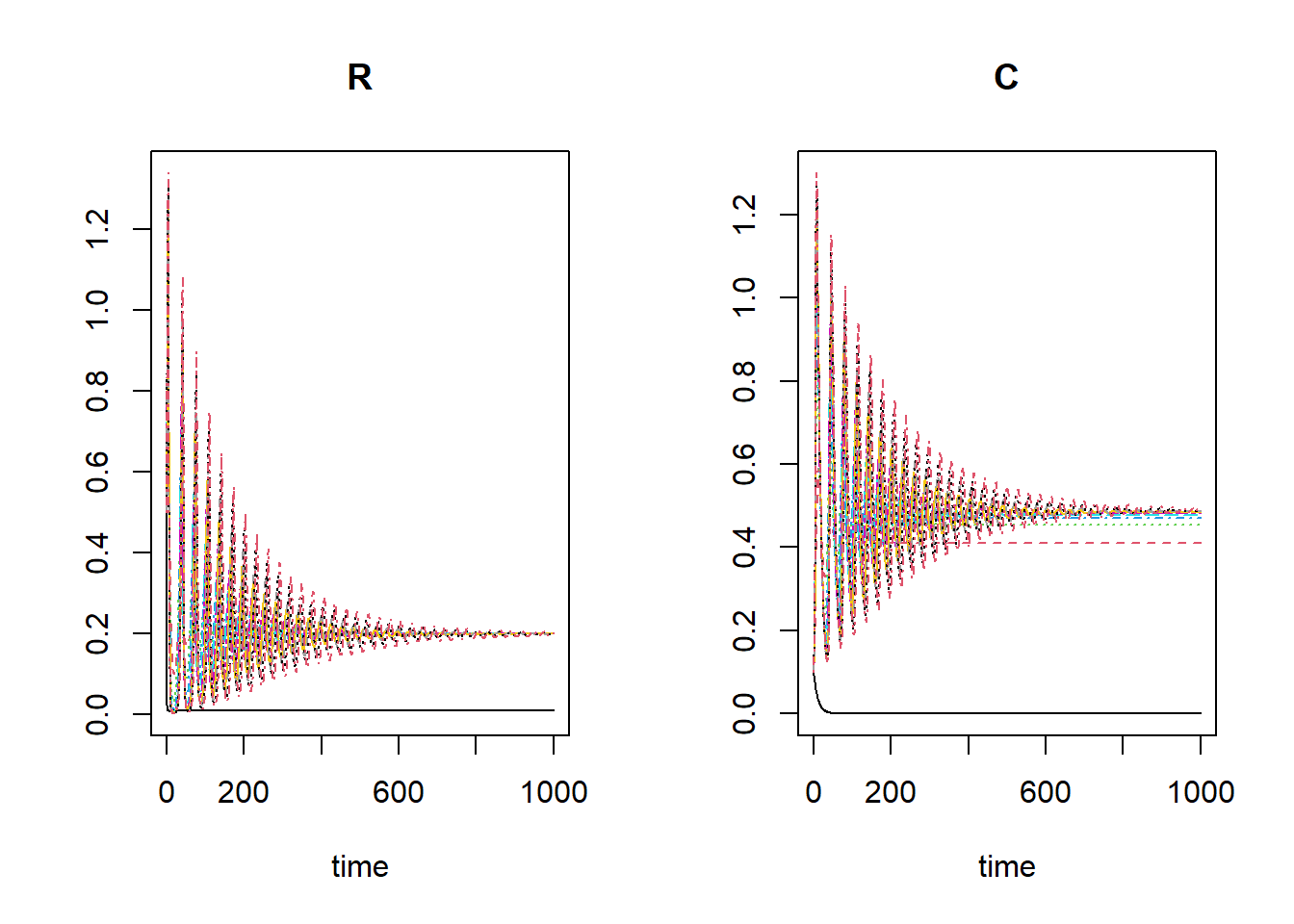
kseq <- seq(from = 0.01, to = 1, length = 10)
outlist <- list()
for (i in 1:length(kseq)) {
outlist[[i]] <- ode(
y = InitPredatorPrey2(),
times = seq(from = 0, to = 1000, by = 1),
func = RatesPredatorPrey2,
parms = ParmsPredatorPrey2(K = kseq[i]),
method = "ode45"
)
}
plot(outlist[[1]], outlist[-1])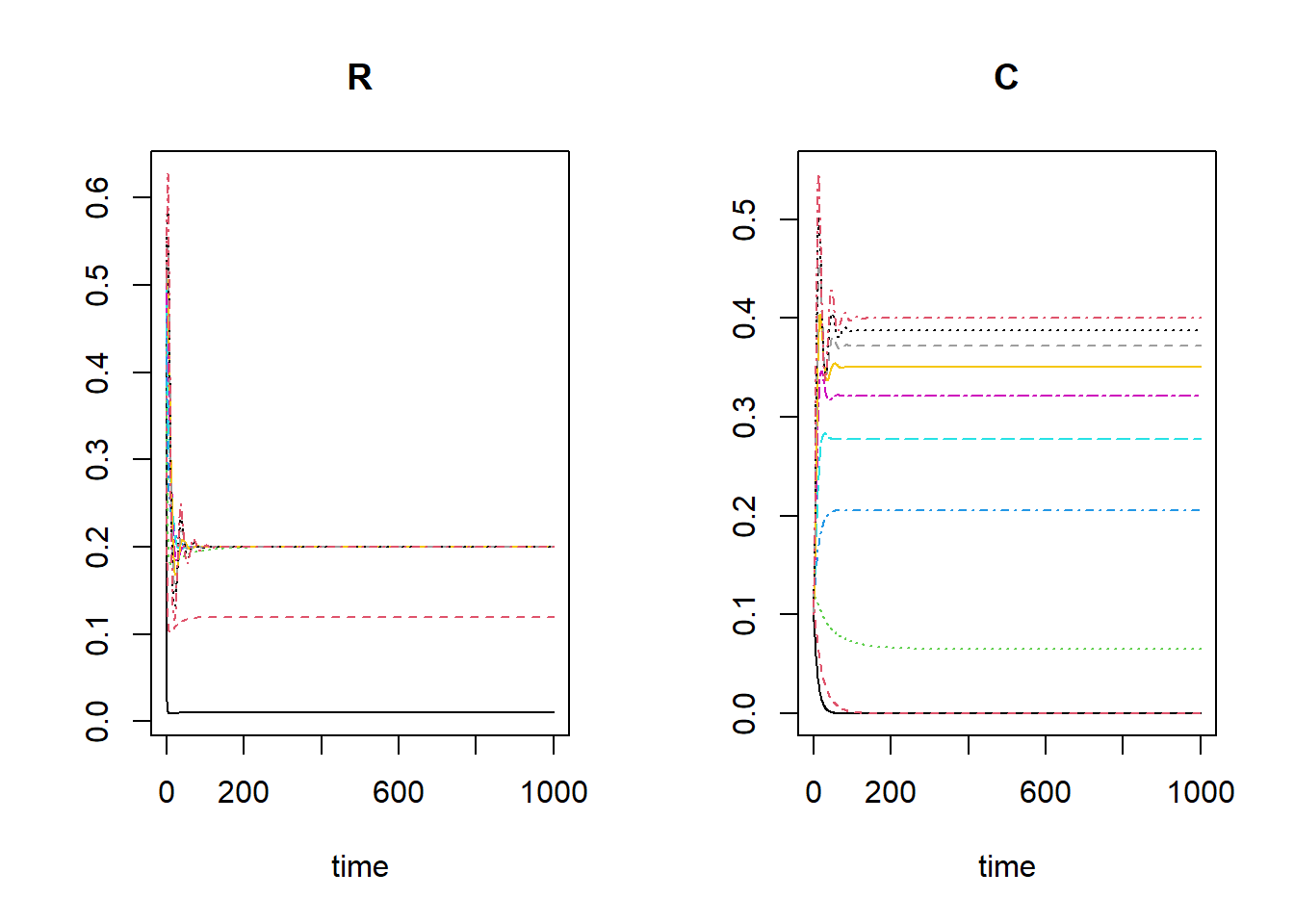
In most scenarios there will be a viable predator populaiton, yet in
some scenarios the predator population will decrease to 0. This is the
case for the ‘r nextinct’ lowest values of kseq: 0.01,
0.12.
Exercise 4.3
The models above assume that the attack/kill rate, , is constant and independent of and . However, this is often not realistic, e.g. when there is saturation in the need for food (e.g. as in the classical Rosenzweig-MacArthur predator-prey model), or when interference between predators reduces predation efficiency. Let’s assume that is not constant, but rather a function of and , as well as the parameters and :
Everything else being equal, more predators will now lead to a lower attack efficiency. Do not bother too much with the ecological mechanism/realism of this equation: we now simply use it to practice skills for numerical integration!
attackRate using
the formula above, with the default values 0.0 for and 0.1 for . Note that the values and result in a model that is actually
the same as in the previous exercise! Plot the attack rate for
R = 1:10, and C = c(0.25, 0.5, 1.0). Plot
separate lines for the three values of , in different colours so that you can
easily identify the scenarios. attackRate <- function(R, C, b1 = 0.0, b2 = 0.1) {
y <- b1 + b2 * R / C
return(y)
}
plot(1:10, attackRate(1:10, 0.25), type = "l", ylim = c(0, 5), xlab = "R", ylab = "attack rate")
lines(1:10, attackRate(1:10, 0.5), col = "red")
lines(1:10, attackRate(1:10, 1), col = "green")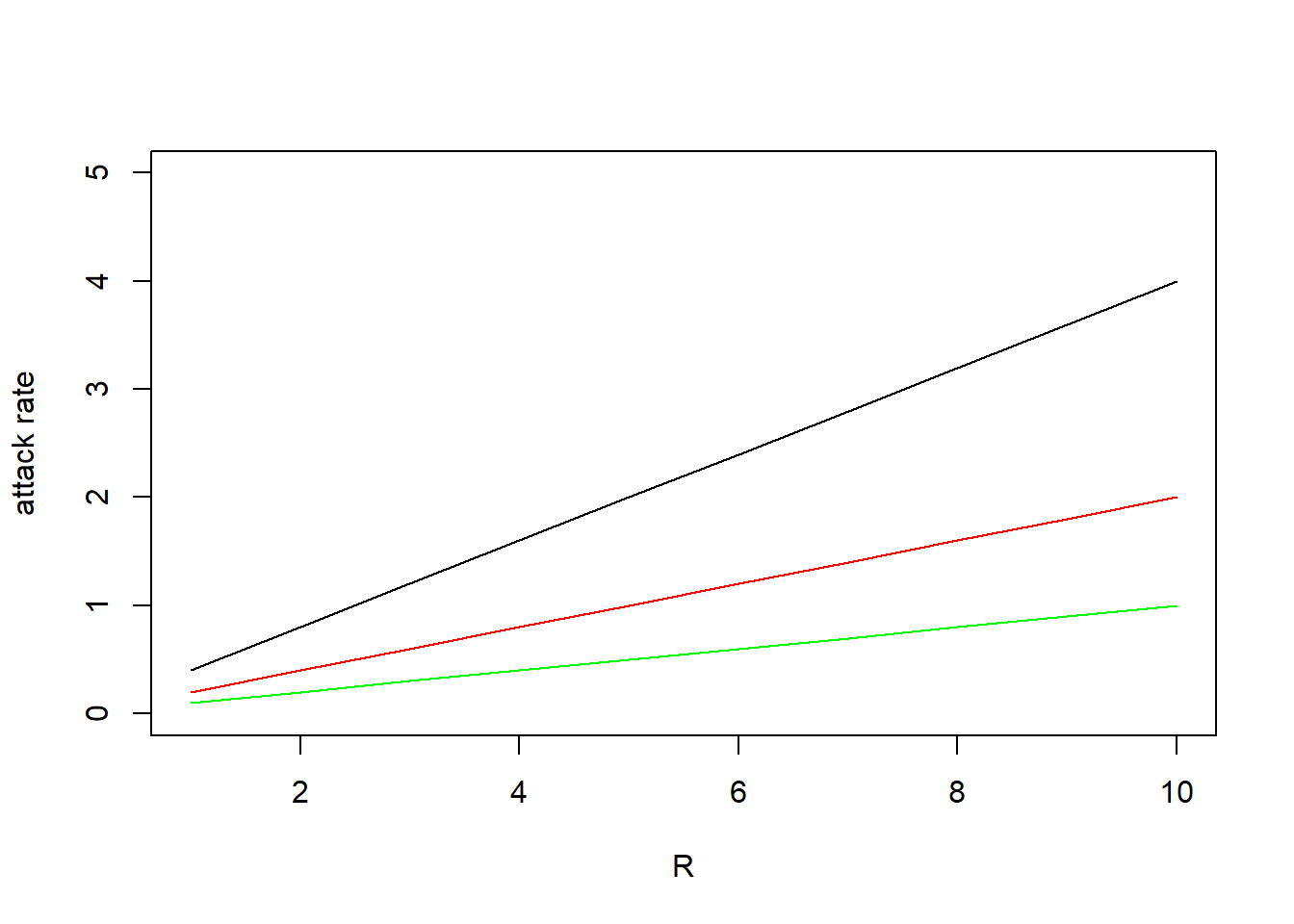
InitPredatorPrey2,
ParmsPredatorPrey2 and RatesPredatorPrey2
functions, and update where necessary! Give these functions the names
InitPredatorPrey3 ParmsPredatorPrey3 and
RatesPredatorPrey3 for this exercise. Note that the attack
rate is used in 2 parts of the model: in the mortality of , and in the growth or . Thus, it may be most efficient to first
compute the attack rate and assign it to an (auxiliary) variable,
e.g. ar, and then use this variable in the rate equations.
An advantage then is that you can choose to return the value of the
auxiliary variable (remember: the first element of the list returned by
RatesPredatorPrey3 contains the rate variables, so
everything but the first element can be used to include
auxiliary variables! See section 4.6 of the syllabus for examples).
InitPredatorPrey3 <- function(R = 0.5, # Resource / prey
C = 0.1){ # Consumer / predator)
# Gather function arguments in a named vector
y <- c(
R = R,
C = C
)
# Return the named vector
return(y)
}
ParmsPredatorPrey3 <- function(r = 0.5, # intrinsic growth rate of logistic growth
K = 10, # carrying capacity
b1 = 1.0, # attack/kill rate intercept
b2 = 0.0, # attack/kill rate slope
c = 0.5, # conversion efficiency
m = 0.1){ # mortality rate of the predators in the absence of prey
# Gather function arguments in a named vector
y <- c(
r = r,
K = K,
b1 = b1,
b2 = b2,
c = c,
m = m
)
# Return the named vector
return(y)
}
# Differences compared to exercise 4.2:
# - auxiliary function added
# - rate equations adjusted
# - 'ar' is returned as additional data
RatesPredatorPrey3 <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# (thus combine 'y' and 'parms' first, using the function c(), then convert to list using as.list())
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)), {
# Optional: forcing functions: get value of the driving variables at time t (if any)
# Optional: auxiliary equations
ar <- attackRate(R, C, b1, b2)
# Rate equations
dRdt <- r * R * (1 - R / K) - ar * R * C
dCdt <- c * ar * R * C - m * C
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(
dRdt = dRdt,
dCdt = dCdt
)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- c(ar = ar)
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(
c(RATES, MB), # the rates of change
AUX
) # additional output per time step
return(outList)
}
)
}
tseq <- seq(from = 0, to = 100, by = 1)Solve the updated model for 3 scenarios:
- b1=0.1, b2=0.0
- b1=0.05, b2=0.05
- b1=0.0, b2=0.01
states1 <- ode(
y = InitPredatorPrey3(),
times = tseq,
func = RatesPredatorPrey3,
parms = ParmsPredatorPrey3(b1 = 0.1, b2 = 0.0),
method = "ode45"
)
states2 <- ode(
y = InitPredatorPrey3(),
times = tseq,
func = RatesPredatorPrey3,
parms = ParmsPredatorPrey3(b1 = 0.05, b2 = 0.05),
method = "ode45"
)
states3 <- ode(
y = InitPredatorPrey3(),
times = tseq,
func = RatesPredatorPrey3,
parms = ParmsPredatorPrey3(b1 = 0.0, b2 = 0.1),
method = "ode45"
)
plot(states1, states2, states3)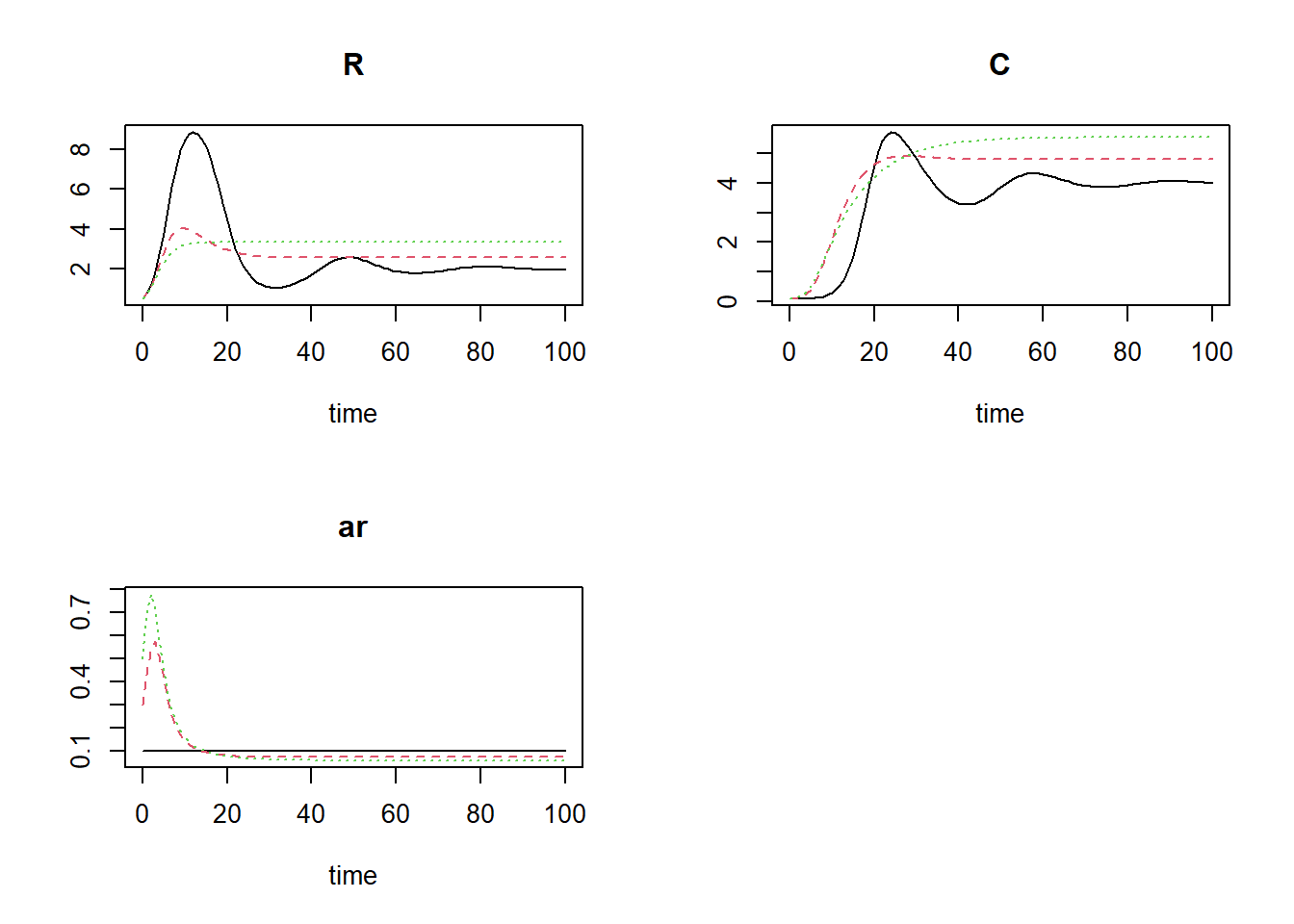
The third scenario (the green line in the plots) will ultimately result in the highest number of prey and predators (yet the attack rate will ultimately be the lowest).
Extra challenges
If you still have time, continue with the exercises below:
Try to find the value of parameter that maximizes the predator density , given the model above with . For this value of , what is the associated attack rate when the system is in balance (i.e. retrieve the value at the end of the integration, after verifying that the simulation has run long enough so that the system is in balance)?
Analyse a new scenario where you use this value (i.e. the associated attack rate) as the value for parameter , where you set parameter . How does the dynamics of this system differ from the one where and is set to the value maximizing the predator density?# create empty list to store output data
outlist <- list()
# specify sequence of values for b2
b2seq <- seq(from = 1e-5, to = 0.1, length.out = 100)
# Iterate over all values of b2, solve model and store output
for(i in seq_along(b2seq)){
outlist[[i]] <- ode(
y = InitPredatorPrey3(R = 1, C = 0.1),
times = seq(from = 0, to = 1000, by = 1),
func = RatesPredatorPrey3,
parms = ParmsPredatorPrey3(b1 = 0.0, b2 = b2seq[[i]]),
method = "ode45"
)
}
# Plot all simulations
plot(outlist[[1]], outlist[-1])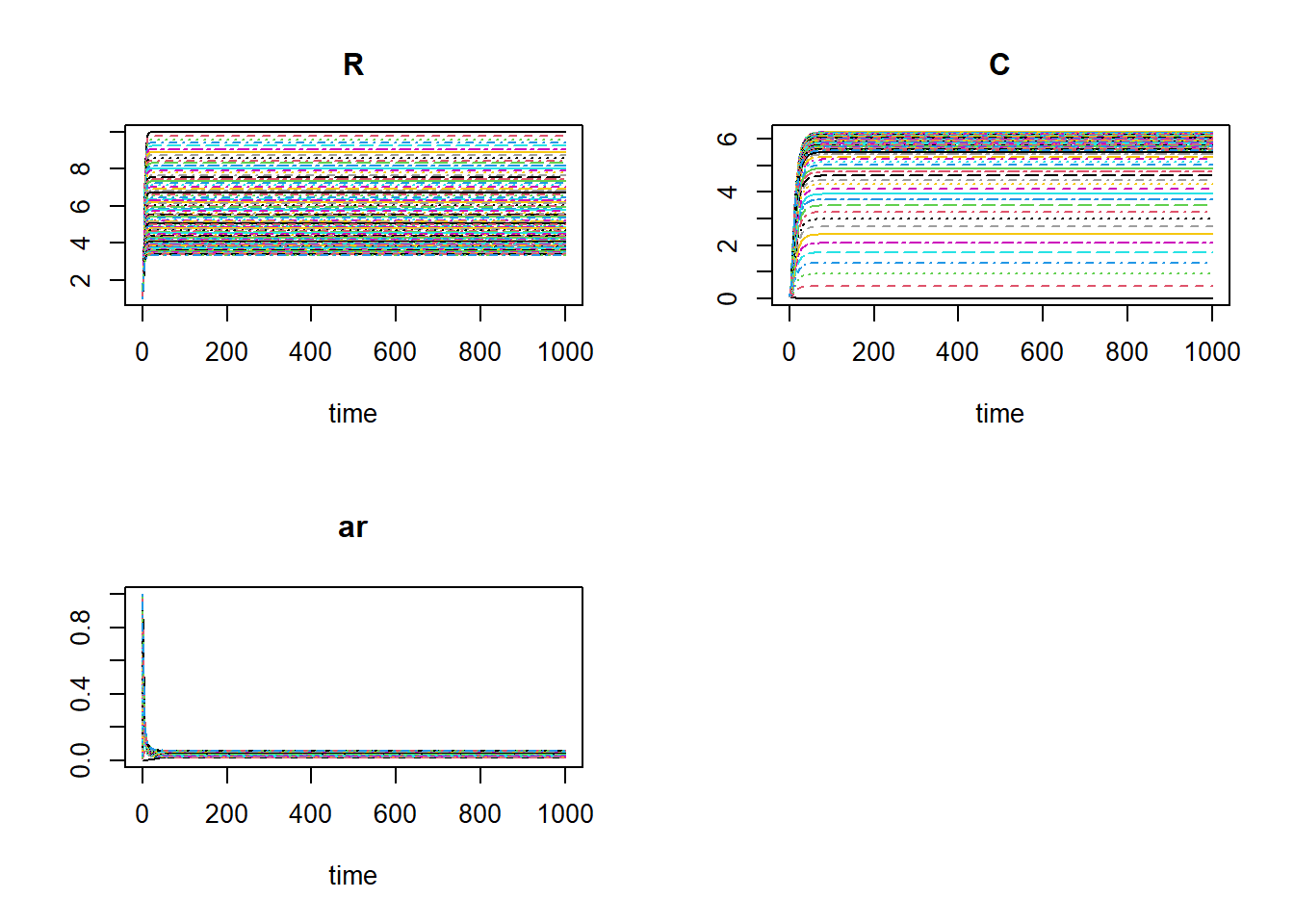
The system seems to have converged in all scenarios.
# Extract (state) values at final timestep
finalValues <- as.data.frame(
do.call(
rbind,
lapply(
outlist,
function(x) {
x[nrow(x), ]
}
)
)
)
head(finalValues)## time R C ar
## 1 1000 9.998000 0.004998001 0.020004
## 2 1000 9.800078 0.489811837 0.020408
## 3 1000 9.609840 0.937342695 0.020812
## 4 1000 9.426848 1.350754944 0.021216
## 5 1000 9.250694 1.732900550 0.021620
## 6 1000 9.081003 2.086354562 0.022024# get value of b2 that maximizes the value for C
maxB2 <- b2seq[which.max(finalValues$C)]
maxB2## [1] 0.0495# Plot (with b2 value maximizing C indicated as vertical dashed line)
par(mfrow = c(2, 2))
plot(b2seq, finalValues$R, type = "l", xlab = "b", ylab = "R")
abline(v = maxB2, col = 8, lty = 2)
plot(b2seq, finalValues$C, type = "l", xlab = "b", ylab = "C")
abline(v = maxB2, col = 8, lty = 2)
plot(b2seq, finalValues$ar, type = "l", xlab = "b", ylab = "attack rate")
abline(v = maxB2, col = 8, lty = 2)
par(mfrow = c(1, 1))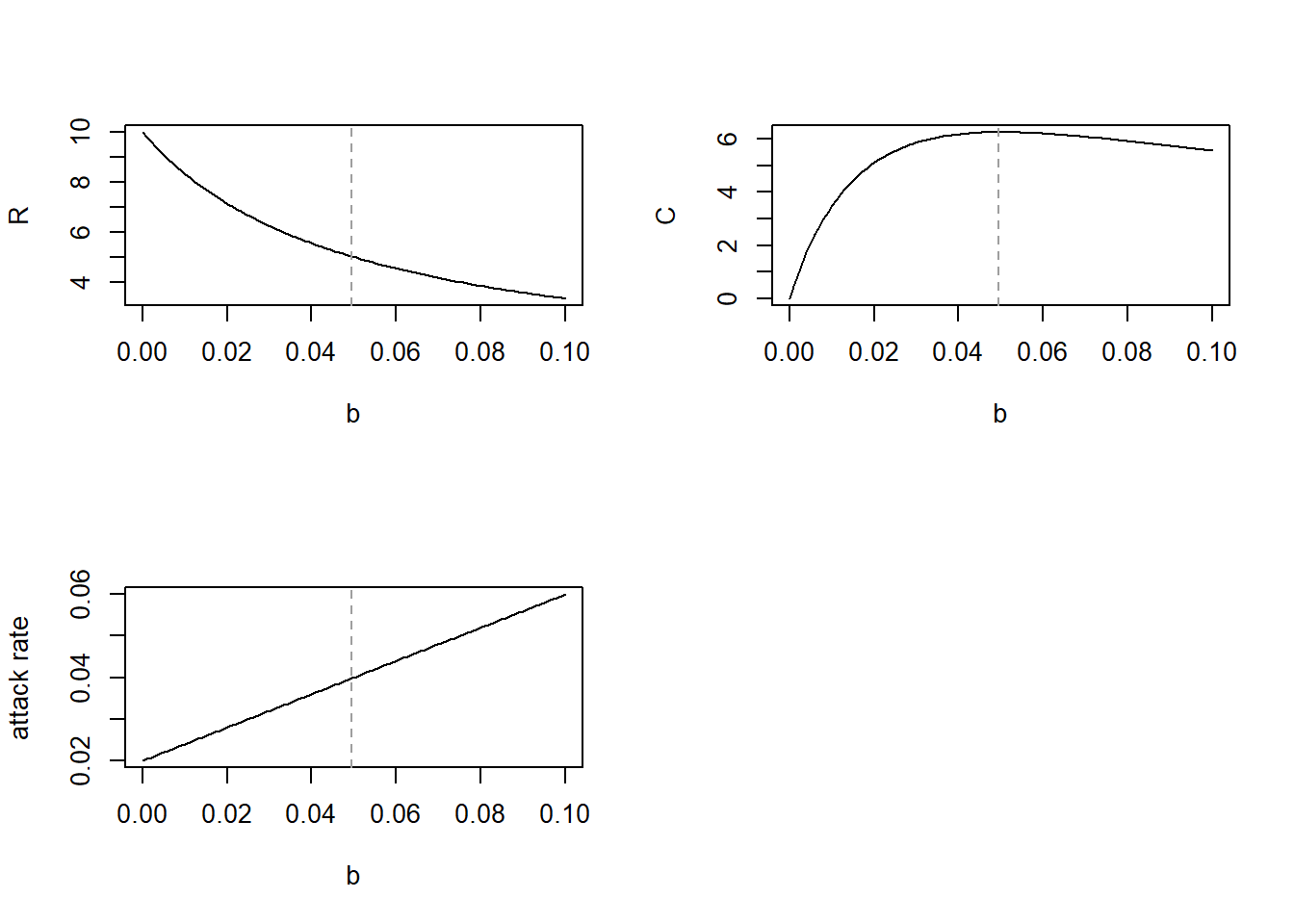
# What is the corresponding ar value at maximum C?
finalValues$ar[which.max(finalValues$C)]## [1] 0.0398# solve model with the b2 value that maximizes C (b1 = 0, b2 = 0.0495)
states1 <- ode(
y = InitPredatorPrey3(R = 1, C = 0.01),
times = seq(from = 0, to = 200, by = 1),
func = RatesPredatorPrey3,
parms = ParmsPredatorPrey3(b1 = 0.0, b2 = 0.0495),
method = "ode45"
)
# solve model with corresponding ar value (b1 = 0.0398, b2 = 0)
states2 <- ode(
y = InitPredatorPrey3(R = 1, C = 0.01),
times = seq(from = 0, to = 200, by = 1),
func = RatesPredatorPrey3,
parms = ParmsPredatorPrey3(b1 = 0.0398, b2 = 0),
method = "ode45"
)
# Plot both
plot(states1, states2)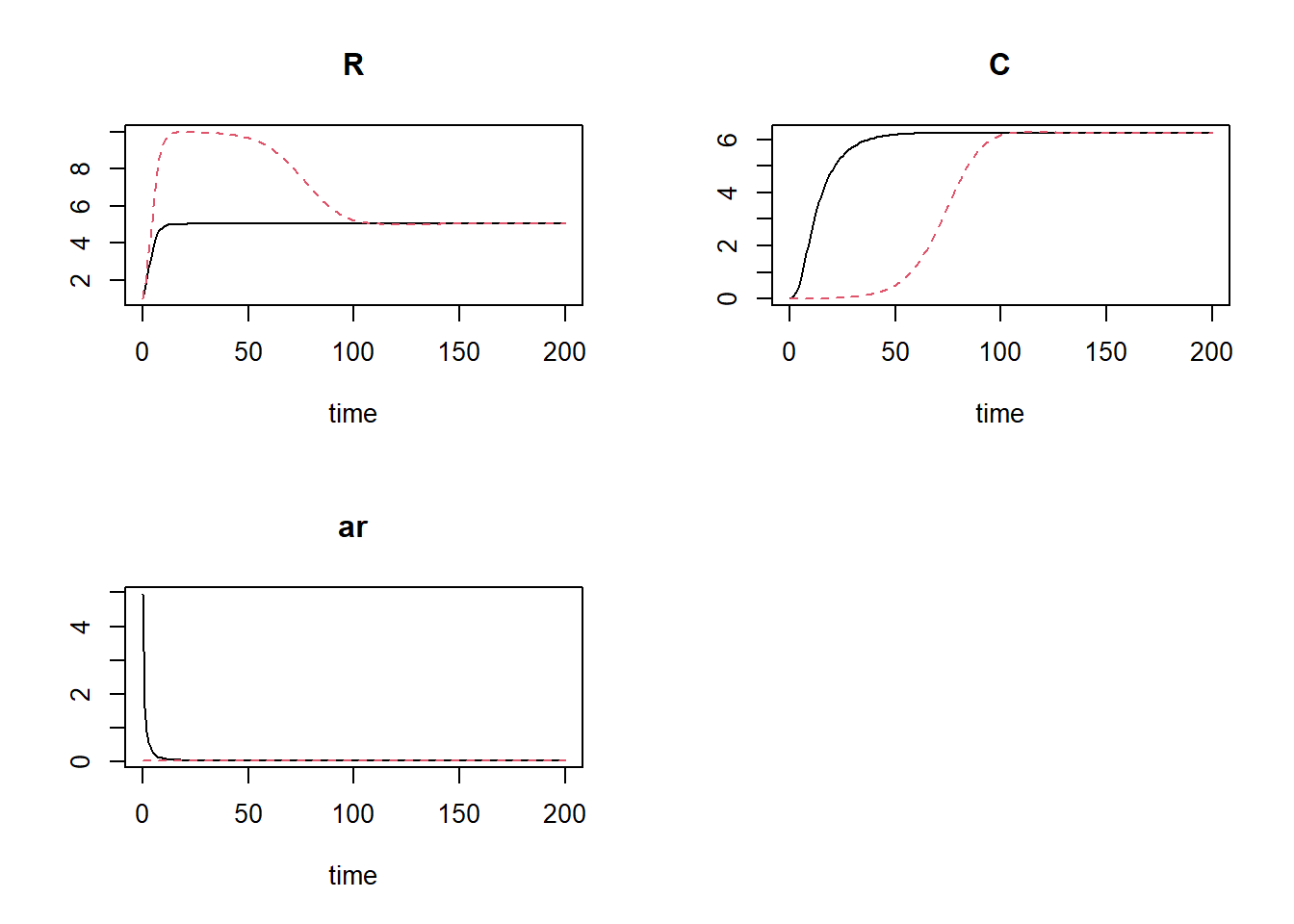
We see that now in both scenarios the system converges to the same equilibrium, yet with the model specification with parameter set to its optimum value (in terms of state ), the system converges much quicker.
forcingK as created on page 94 of the
syllabus, as well as the information on that page, to include not as a constant, but a forcing
function. Note: in the models in these exercises we’ve set to 10, wereas the forcingK
data is on average ca 100, thus divide the forcingK data
(the column ) as retrieved from page
94 by 10 to have a time series that is variable yet on average equal to
the value used in the previous exercises. The time series covers 50 time
units, thus solve the ODE with as a
forcing function for 50 time units.
Template 2 (on
driving variables) can be of use here. # data frame with info on driving variable
forcingK <- cbind(
0:50,
c(
84, 82, 53, 67, 74, 123, 150, 136, 92, 105, 120, 136, 144, 117, 120, 136, 125,
93, 108, 123, 113, 149, 112, 112, 51, 64, 109, 112, 98, 115, 90, 102, 65, 50,
63, 66, 125, 150, 121, 120, 125, 120, 121, 139, 113, 96, 115, 82, 97, 101, 85
)
)
# create forcing function
Kfun <- approxfun(x = forcingK[,1], y = forcingK[,2] / 10, method = "linear", rule = 2)
# update parameters function, remove K
ParmsPredatorPrey4 <- function(r = 0.5, # intrinsic growth rate of logistic growth
b1 = 1.0, # attack/kill rate intercept
b2 = 0.0, # attack/kill rate slope
c = 0.5, # conversion efficiency
m = 0.1){ # mortality rate of the predators in the absence of prey
# Gather function arguments in a named vector
y <- c(
r = r,
b1 = b1,
b2 = b2,
c = c,
m = m
)
# Return the named vector
return(y)
}
# update function with rate equations
# - difference with version 2 above: added forcing function and added as output to return list
RatesPredatorPrey4 <- function(t, y, parms) {
# use the with() function to be able to access the parameters and states easily
# remember that the with() function is with(data, expr), where
# - 'data' is ideally a list with named elements
# (thus combine 'y' and 'parms' first, using the function c(), then convert to list using as.list())
# - 'expr' is an expression (i.e. code) that is evaluated
# this can span multiple lines of code when embraced with curly brackets {}
with(
as.list(c(y, parms)), {
# Optional: forcing functions: get value of the driving variables at time t (if any)
K <- Kfun(t)
# Optional: auxiliary equations
ar <- attackRate(R, C, b1, b2)
# Rate equations
dRdt <- r * R * (1 - R / K) - ar * R * C
dCdt <- c * ar * R * C - m * C
# Gather all rates of change in a vector
# - the rates should be in the same order as the states (as specified in 'y')
# - it can be a named vector, but does not need to be
RATES <- c(
dRdt = dRdt,
dCdt = dCdt
)
# Optional: get in/out flow used to compute mass balances (or set MB <- NULL)
MB <- NULL
# Optional: gather auxiliary variables that should be returned (or set AUX <- NULL)
# - this should be a named vector or list!
AUX <- c(
ar = ar,
K = K
)
# Return result as a list
# - the first element is a vector with the rates of change (in the same order as 'y')
# - all other elements are (optional) extra output, which should be named
outList <- list(
c(RATES, MB), # the rates of change
AUX
) # additional output per time step
return(outList)
}
)
}
# solve
states <- ode(
y = InitPredatorPrey3(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesPredatorPrey4,
parms = ParmsPredatorPrey4(),
method = "ode45"
)
head(states)## time R C ar K
## [1,] 0.0 0.5000000 0.1000000 1 8.40
## [2,] 0.1 0.5187882 0.1015588 1 8.38
## [3,] 0.2 0.5381300 0.1032402 1 8.36
## [4,] 0.3 0.5580257 0.1050525 1 8.34
## [5,] 0.4 0.5784735 0.1070044 1 8.32
## [6,] 0.5 0.5994695 0.1091056 1 8.30plot(states)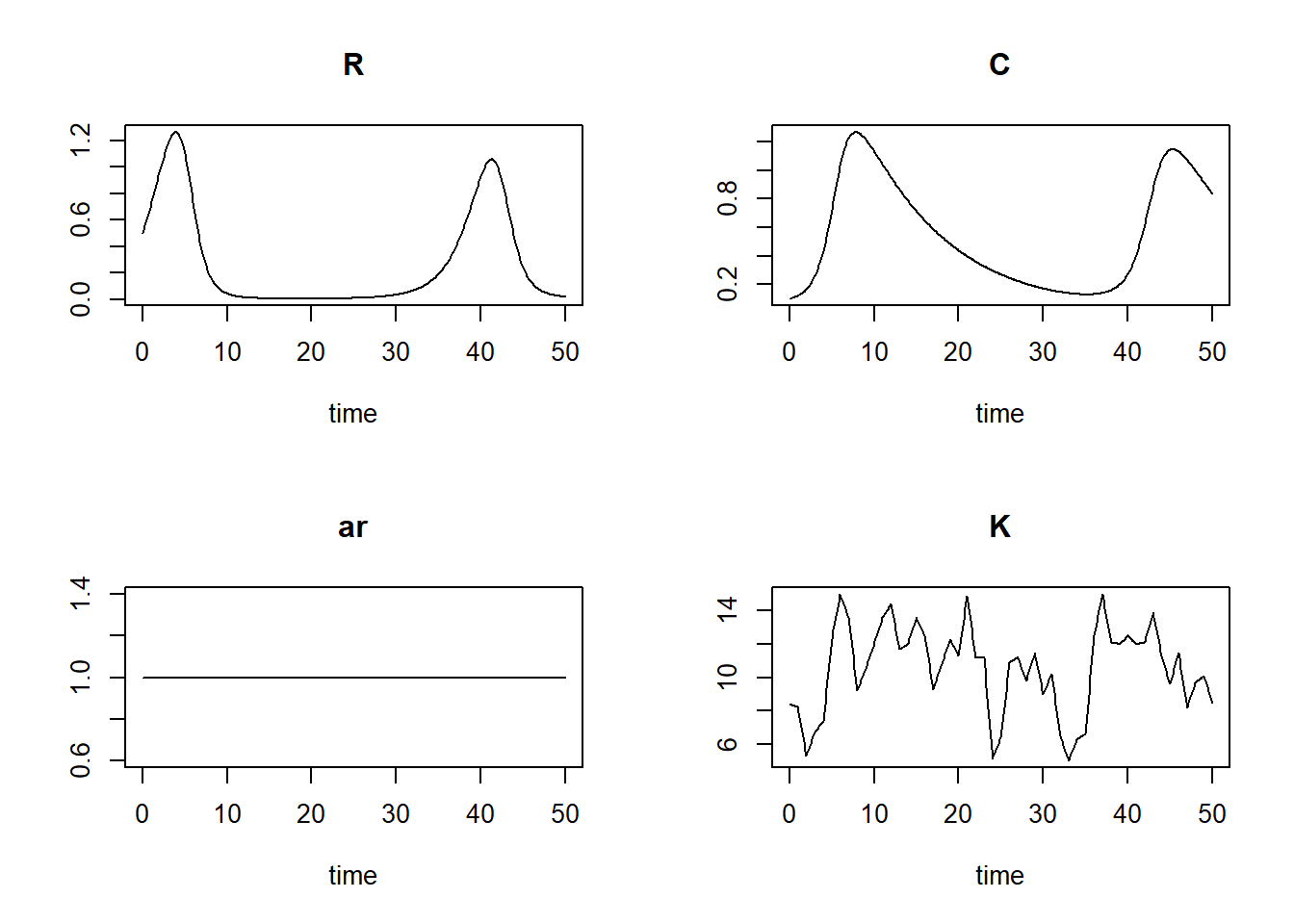
# Define events
events <- list(data = data.frame(
var = "R",
time = c(10, 20, 30, 40),
value = 0.75,
method = "multiply"
))
# solve including events
states <- ode(
y = InitPredatorPrey3(),
times = seq(from = 0, to = 50, by = 0.1),
func = RatesPredatorPrey4,
parms = ParmsPredatorPrey4(),
method = "ode45",
events = events
)
# plot
plot(states)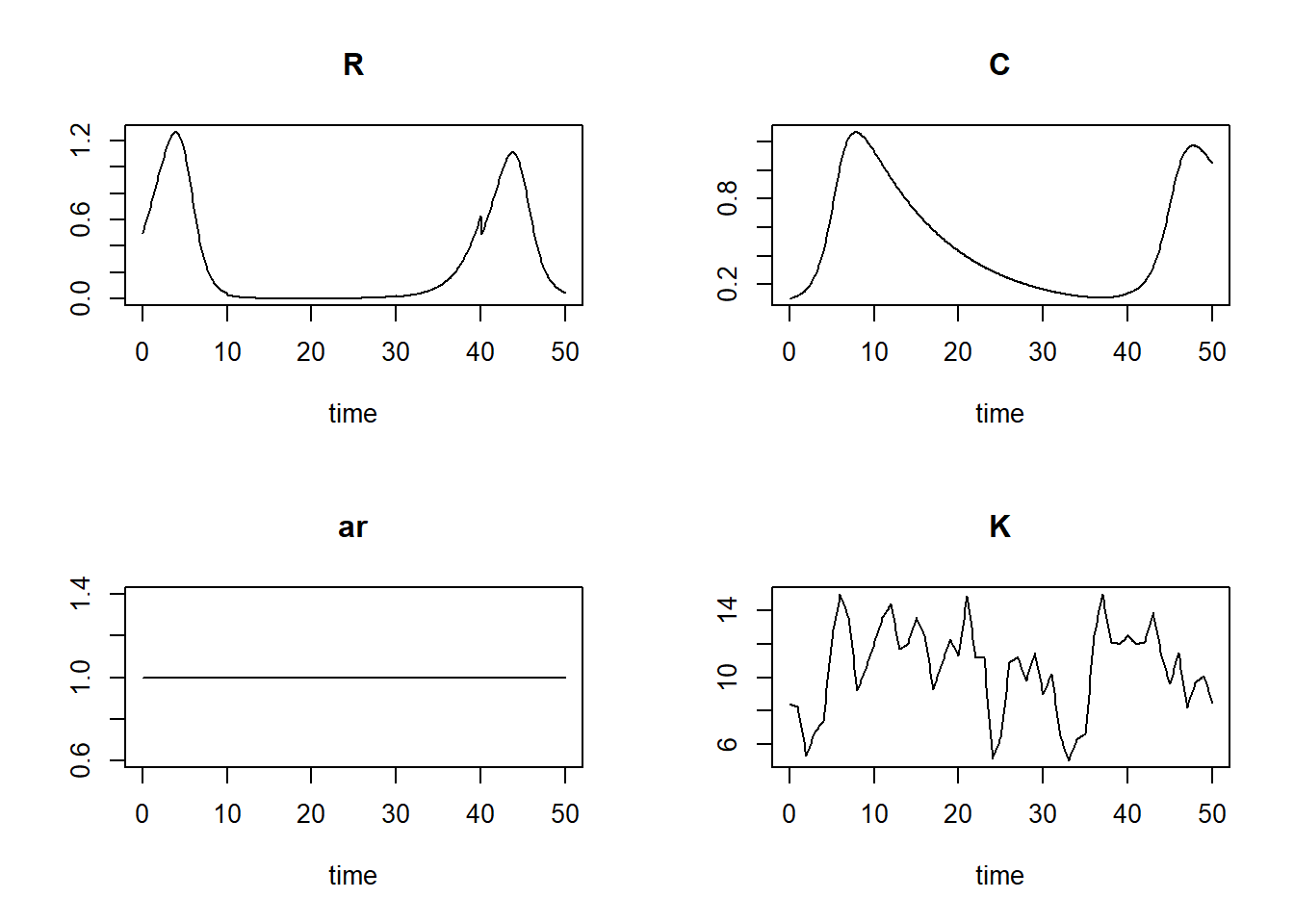
Solutions as R script
Download here the code as shown on this page in a separate .r file.